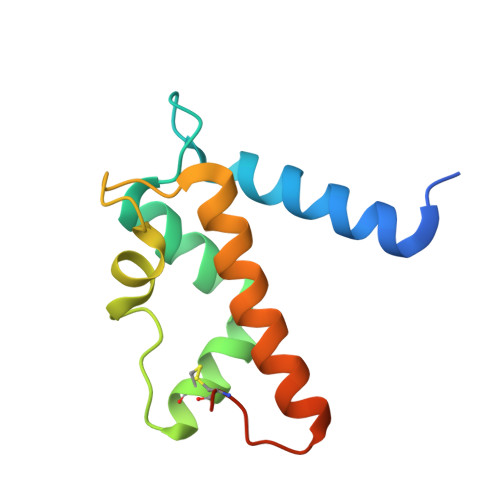
Structural Characterization of S100A15 Reveals a Novel Zinc Coordination Site Among S100 Proteins and Altered Surface Chemistry with Functional Implications for Receptor Binding.
Murray, J.I., Tonkin, M.L., Whiting, A.L., Peng, F., Farnell, B., Cullen, J.T., Hof, F., Boulanger, M.J.(2012) BMC Struct Biol 12: 16
- PubMed: 22747601
- DOI: https://doi.org/10.1186/1472-6807-12-16
- Primary Citation of Related Structures:
4AQI, 4AQJ - PubMed Abstract:
S100 proteins are a family of small, EF-hand containing calcium-binding signaling proteins that are implicated in many cancers. While the majority of human S100 proteins share 25-65% sequence similarity, S100A7 and its recently identified paralog, S100A15, display 93% sequence identity. Intriguingly, however, S100A7 and S100A15 serve distinct roles in inflammatory skin disease; S100A7 signals through the receptor for advanced glycation products (RAGE) in a zinc-dependent manner, while S100A15 signals through a yet unidentified G-protein coupled receptor in a zinc-independent manner. Of the seven divergent residues that differentiate S100A7 and S100A15, four cluster in a zinc-binding region and the remaining three localize to a predicted receptor-binding surface.
Organizational Affiliation:
Department of Chemistry, University of Victoria, PO Box 3065STN CSC, Victoria, BC V8W 3P6, Canada.