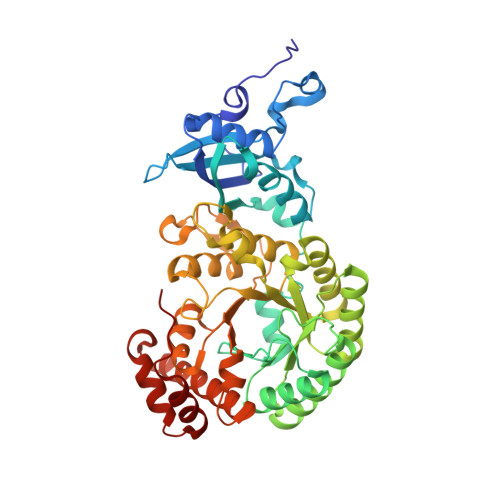
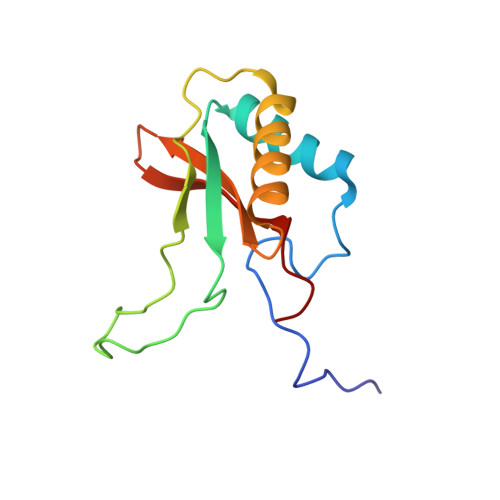
Identification of Interactions between Abscisic Acid and Ribulose-1,5-Bisphosphate Carboxylase/Oxygenase.
Galka, M.M., Rajagopalan, N., Buhrow, L.M., Nelson, K.M., Switala, J., Cutler, A.J., Palmer, D.R., Loewen, P.C., Abrams, S.R., Loewen, M.C.(2015) PLoS One 10: e0133033-e0133033
- PubMed: 26197050
- DOI: https://doi.org/10.1371/journal.pone.0133033
- Primary Citation of Related Structures:
4MKV - PubMed Abstract:
Abscisic acid ((+)-ABA) is a phytohormone involved in the modulation of developmental processes and stress responses in plants. A chemical proteomics approach using an ABA mimetic probe was combined with in vitro assays, isothermal titration calorimetry (ITC), x-ray crystallography and in silico modelling to identify putative (+)-ABA binding-proteins in crude extracts of Arabidopsis thaliana. Ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) was identified as a putative ABA-binding protein. Radiolabelled-binding assays yielded a Kd of 47 nM for (+)-ABA binding to spinach Rubisco, which was validated by ITC, and found to be similar to reported and experimentally derived values for the native ribulose-1,5-bisphosphate (RuBP) substrate. Functionally, (+)-ABA caused only weak inhibition of Rubisco catalytic activity (Ki of 2.1 mM), but more potent inhibition of Rubisco activation (Ki of ~ 130 μM). Comparative structural analysis of Rubisco in the presence of (+)-ABA with RuBP in the active site revealed only a putative low occupancy (+)-ABA binding site on the surface of the large subunit at a location distal from the active site. However, subtle distortions in electron density in the binding pocket and in silico docking support the possibility of a higher affinity (+)-ABA binding site in the RuBP binding pocket. Overall we conclude that (+)-ABA interacts with Rubisco. While the low occupancy (+)-ABA binding site and weak non-competitive inhibition of catalysis may not be relevant, the high affinity site may allow ABA to act as a negative effector of Rubisco activation.
Organizational Affiliation:
National Research Council of Canada, Saskatoon, Saskatchewan, Canada; Department of Chemistry, University of Saskatchewan, Saskatoon, Saskatchewan, Canada.