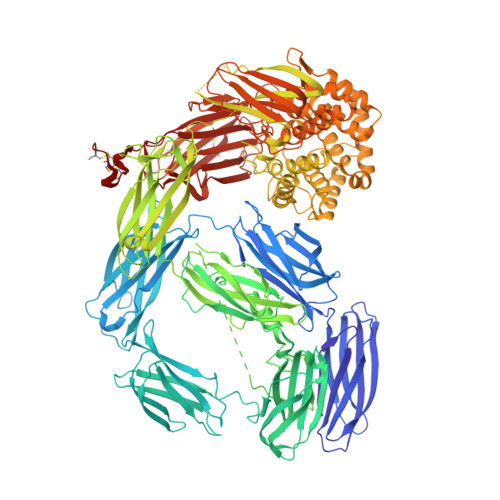
Molecular basis for genetic resistance of Anopheles gambiae to Plasmodium: structural analysis of TEP1 susceptible and resistant alleles.
Le, B.V., Williams, M., Logarajah, S., Baxter, R.H.(2012) PLoS Pathog 8: e1002958-e1002958
- PubMed: 23055931
- DOI: https://doi.org/10.1371/journal.ppat.1002958
- Primary Citation of Related Structures:
4D94, 4LNV - PubMed Abstract:
Thioester-containing protein 1 (TEP1) is a central component in the innate immune response of Anopheles gambiae to Plasmodium infection. Two classes of TEP1 alleles, TEP1*S and TEP1*R, are found in both laboratory strains and wild isolates, related by a greater or lesser susceptibility, respectively to both P. berghei and P. falciparum infection. We report the crystal structure of the full-length TEP1*S1 allele which, while similar to the previously determined structure of full-length TEP1*R1, displays flexibility in the N-terminal fragment comprising domains MG1-MG6. Amino acid differences between TEP1*R1 and TEP1*S1 are localized to the TED-MG8 domain interface that protects the thioester bond from hydrolysis and structural changes are apparent at this interface. As a consequence cleaved TEP1*S1 (TEP1*S1(cut)) is significantly more susceptible to hydrolysis of its intramolecular thioester bond than TEP1*R1(cut). TEP1*S1(cut) is stabilized in solution by the heterodimeric LRIM1/APL1C complex, which preserves the thioester bond within TEP1*S1(cut). These results suggest a mechanism by which selective pressure on the TEP1 gene results in functional variation that may influence the vector competence of A. gambiae towards Plasmodium infection.
Organizational Affiliation:
Department of Chemistry and Molecular Biophysics & Biochemistry, Yale University, New Haven, Connecticut, United States of America.