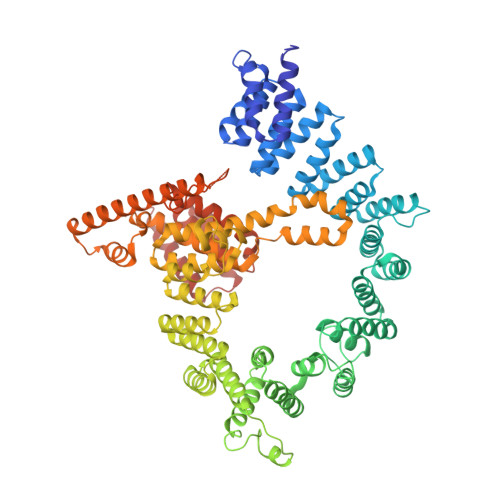
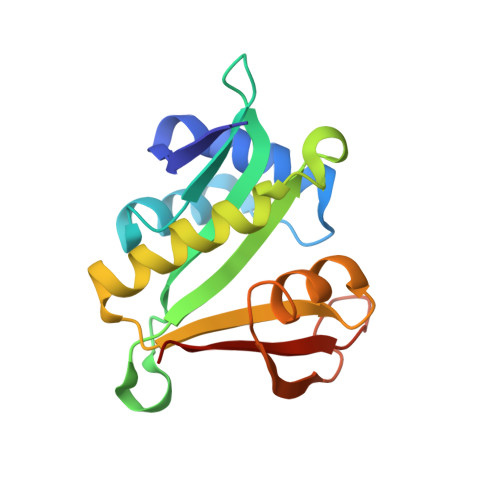
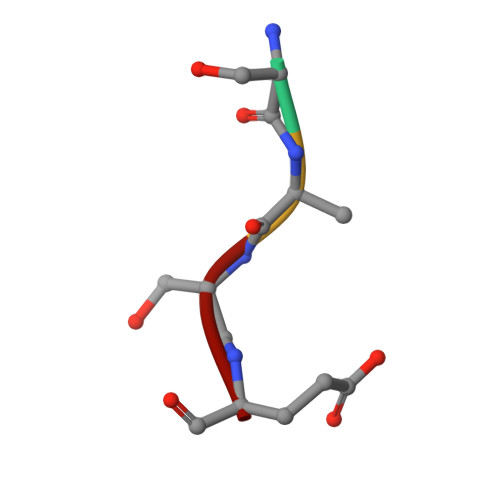
Molecular basis for N-terminal acetylation by the heterodimeric NatA complex.
Liszczak, G., Goldberg, J.M., Foyn, H., Petersson, E.J., Arnesen, T., Marmorstein, R.(2013) Nat Struct Mol Biol 20: 1098-1105
- PubMed: 23912279
- DOI: https://doi.org/10.1038/nsmb.2636
- Primary Citation of Related Structures:
4KVM, 4KVO, 4KVX - PubMed Abstract:
N-terminal acetylation is ubiquitous among eukaryotic proteins and controls a myriad of biological processes. Of the N-terminal acetyltransferases (NATs) that facilitate this cotranslational modification, the heterodimeric NatA complex has the most diversity for substrate selection and modifies the majority of all N-terminally acetylated proteins. Here, we report the X-ray crystal structure of the 100-kDa holo-NatA complex from Schizosaccharomyces pombe, in the absence and presence of a bisubstrate peptide-CoA-conjugate inhibitor, as well as the structure of the uncomplexed Naa10p catalytic subunit. The NatA-Naa15p auxiliary subunit contains 13 tetratricopeptide motifs and adopts a ring-like topology that wraps around the NatA-Naa10p subunit, an interaction that alters the Naa10p active site for substrate-specific acetylation. These studies have implications for understanding the mechanistic details of other NAT complexes and how regulatory subunits modulate the activity of the broader family of protein acetyltransferases.
Organizational Affiliation:
1] Program in Gene Expression and Regulation, Wistar Institute, Philadelphia, Pennsylvania, USA. [2] Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania, USA.