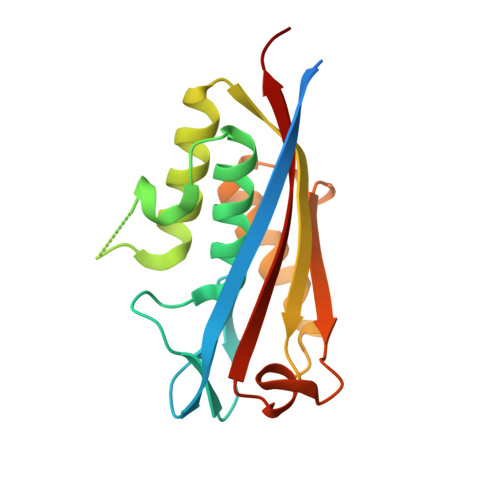
Crystal Structures of Ispf from Plasmodium Falciparum and Burkholderia Cenocepacia: Comparisons Inform Antimicrobial Drug Target Assessment.
O'Rourke, P.E.F., Kalinowska-Tluscik, J., Fyfe, P.K., Dawson, A., Hunter, W.N.(2014) BMC Struct Biol 14: 1
- PubMed: 24410837
- DOI: https://doi.org/10.1186/1472-6807-14-1
- Primary Citation of Related Structures:
4C81, 4C82, 4C8E, 4C8G, 4C8I - PubMed Abstract:
2C-methyl-D-erythritol-2,4-cyclodiphosphate synthase (IspF) catalyzes the conversion of 4-diphosphocytidyl-2C-methyl-D-erythritol-2-phosphate to 2C-methyl-D-erythritol-2,4-cyclodiphosphate and cytidine monophosphate in production of isoprenoid-precursors via the methylerythritol phosphate biosynthetic pathway. IspF is found in the protozoan Plasmodium falciparum, a parasite that causes cerebral malaria, as well as in many Gram-negative bacteria such as Burkholderia cenocepacia. IspF represents a potential target for development of broad-spectrum antimicrobial drugs since it is proven or inferred as essential in these pathogens and absent from mammals. Structural studies of IspF from these two important yet distinct pathogens, and comparisons with orthologues have been carried out to generate reagents, to support and inform a structure-based approach to early stage drug discovery.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee, DD1 5EH, UK. w.n.hunter@dundee.ac.uk.