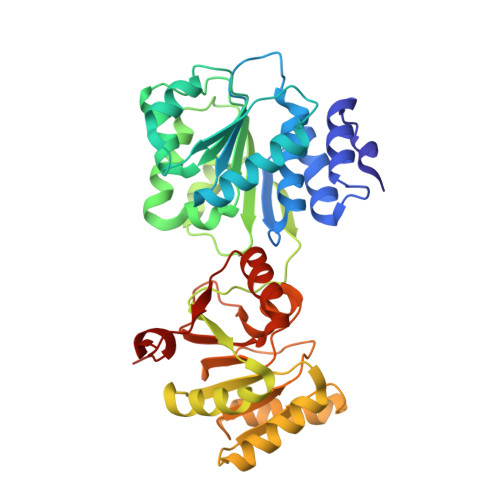
Structural Analysis of the Yeast Dhh1-Pat1 Complex Reveals How Dhh1 Engages Pat1, Edc3 and RNA in Mutually Exclusive Interactions
Sharif, H., Ozgur, S., Sharma, K., Basquin, C., Urlaub, H., Conti, E.(2013) Nucleic Acids Res 41: 8377
- PubMed: 23851565
- DOI: https://doi.org/10.1093/nar/gkt600
- Primary Citation of Related Structures:
4BRU, 4BRW - PubMed Abstract:
Translational repression and deadenylation of eukaryotic mRNAs result either in the sequestration of the transcripts in a nontranslatable pool or in their degradation. Removal of the 5' cap structure is a crucial step that commits deadenylated mRNAs to 5'-to-3' degradation. Pat1, Edc3 and the DEAD-box protein Dhh1 are evolutionary conserved factors known to participate in both translational repression and decapping, but their interplay is currently unclear. We report the 2.8 Å resolution structure of yeast Dhh1 bound to the N-terminal domain of Pat1. The structure shows how Pat1 wraps around the C-terminal RecA domain of Dhh1, docking onto the Phe-Asp-Phe (FDF) binding site. The FDF-binding site of Dhh1 also recognizes Edc3, revealing why the binding of Pat1 and Edc3 on Dhh1 are mutually exclusive events. Using co-immunoprecipitation assays and structure-based mutants, we demonstrate that the mode of Dhh1-Pat1 recognition is conserved in humans. Pat1 and Edc3 also interfere and compete with the RNA-binding properties of Dhh1. Mapping the RNA-binding sites on Dhh1 with a crosslinking-mass spectrometry approach shows a large RNA-binding surface around the C-terminal RecA domain, including the FDF-binding pocket. The results suggest a model for how Dhh1-containing messenger ribonucleoprotein particles might be remodeled upon Pat1 and Edc3 binding.
Organizational Affiliation:
Structural Cell Biology Department, Max Planck Institute of Biochemistry, Am Klopferspitz 18, Martinsried/Munich, D-82152 Germany and Cellular Biochemistry Department, Max Planck Institute of Biophysical Chemistry, Am Faßberg 11, 37077 Göttingen, Germany.