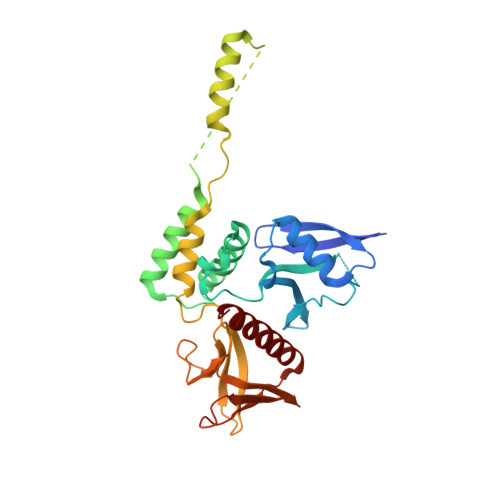
Unfurling of the band 4.1, ezrin, radixin, moesin (FERM) domain of the merlin tumor suppressor.
Yogesha, S.D., Sharff, A.J., Giovannini, M., Bricogne, G., Izard, T.(2011) Protein Sci 20: 2113-2120
- PubMed: 22012890
- DOI: https://doi.org/10.1002/pro.751
- Primary Citation of Related Structures:
3U8Z - PubMed Abstract:
The merlin-1 tumor suppressor is encoded by the Neurofibromatosis-2 (Nf2) gene and loss-of-function Nf2 mutations lead to nervous system tumors in man and to several tumor types in mice. Merlin is an ERM (ezrin, radixin, moesin) family cytoskeletal protein that interacts with other ERM proteins and with components of cell-cell adherens junctions (AJs). Merlin stabilizes the links of AJs to the actin cytoskeleton. Thus, its loss destabilizes AJs, promoting cell migration and invasion, which in Nf2(+/-) mice leads to highly metastatic tumors. Paradoxically, the "closed" conformation of merlin-1, where its N-terminal four-point-one, ezrin, radixin, moesin (FERM) domain binds to its C-terminal tail domain, directs its tumor suppressor functions. Here we report the crystal structure of the human merlin-1 head domain when crystallized in the presence of its tail domain. Remarkably, unlike other ERM head-tail interactions, this structure suggests that binding of the tail provokes dimerization and dynamic movement and unfurling of the F2 motif of the FERM domain. We conclude the "closed" tumor suppressor conformer of merlin-1 is in fact an "open" dimer whose functions are disabled by Nf2 mutations that disrupt this architecture.
Organizational Affiliation:
Cell Adhesion Laboratory, Department of Cancer Biology, The Scripps Research Institute, Jupiter, Florida 33458, USA.