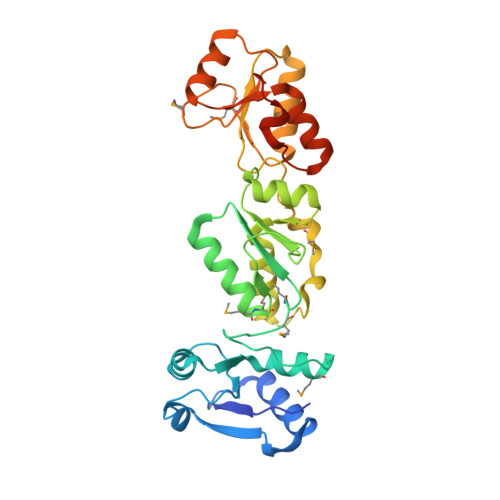
Crystal structure of the N-terminal region of human Topoisomerase II beta binding protein 1
Huo, Y.G., Bai, L., Xu, M., Jiang, T.(2010) Biochem Biophys Res Commun 401: 401-405
- PubMed: 20858457
- DOI: https://doi.org/10.1016/j.bbrc.2010.09.066
- Primary Citation of Related Structures:
3OLC - PubMed Abstract:
Human DNA Topoisomerase IIβ binding protein 1 (TopBP1) is a modulating protein that plays an essential role in the response to DNA damage. The N-terminal region of TopBP1, which contains predicted BRCA1-carboxy terminal (BRCT) domains 1 and 2, binds to Rad9, a component of the cell cycle checkpoint clamp Rad9-Hus1-Rad1 complex. Here, we report the crystal structure of the TopBP1N-terminal region (residues 1-290) at 2.4Å resolution. Interestingly, in addition to the predicted tandem BRCT1-2 repeats (residues 103-284), residues 7-98 form a previously unreported BRCT domain (here, BRCT0). In contrast to both BRCT1 and BRCT2, which possess the conventional phosphopeptide binding residues within a surface pocket, the corresponding pocket in BRCT0 is largely hydrophobic. Structural comparisons together with peptide binding studies indicate that the tandem BRCT1-2 domains are the binding region for phosphorylated Ser387 in Rad9.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, China.