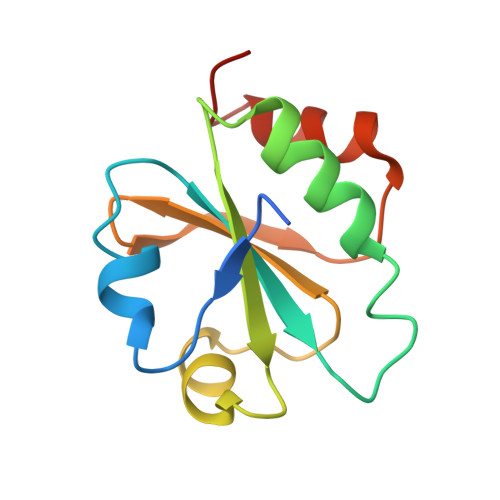
Structure of Mycobacterium tuberculosis thioredoxin in complex with quinol inhibitor PMX464
Hall, G., Bradshaw, T.D., Laughton, C.A., Stevens, M.F., Emsley, J.(2011) Protein Sci 20: 210-215
- PubMed: 20981751
- DOI: https://doi.org/10.1002/pro.533
- Primary Citation of Related Structures:
3NOF, 3O6T - PubMed Abstract:
Thioredoxin (Trx) plays a critical role in the regulation of cellular redox homeostasis. Many disease causing pathogens rely on the Trx redox system for survival in conditions of environmental stress. The Trx redox system has been implicated in the resistance of Mycobacterium tuberculosis (Mtb) to phagocytosis. Trx is able to reduce a variety of target substrates and reactive oxygen species (ROS) through the cyclization of its active site dithiol to the oxidized disulphide Cys37-Cys40. Here we report the crystal structure of the Mtb Trx C active site mutant C40S (MtbTrxCC40S) in isolation and in complex with the hydroxycyclohexadienone inhibitor PMX464. We observe PMX464 is covalently bound to the active site residue Cys37 through Michael addition of the cyclohexadienone ring and also forms noncovalent contacts which mimic the binding of natural Trx ligands. In comparison with the ligand free MtbTrxCC40S structure a conformational change occurs in the PMX464 complex involving movement of helix α2 and the active site loop. These changes are almost identical to those observed for helix α2 in human Trx ligand complexes. Whereas the ligand free structure forms a homodimer the inhibitor complex unexpectedly forms a different dimer with one PMX464 molecule bound at the interface. This 2:1 MtbTrxCC40S-PMX464 complex is also observed using mass spectrometry measurements. This structure provides an unexpected scaffold for the design of improved Trx inhibitors targeted at developing treatments for tuberculosis.
Organizational Affiliation:
Centre for Biomolecular Sciences, School of Pharmacy, University of Nottingham, Nottingham NG7 2RD, United Kingdom.