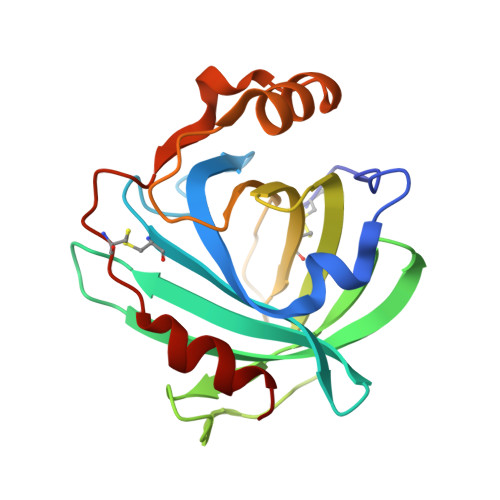
Effect of mutation of carboxyl side-chain amino acids near the heme on the midpoint potentials and ligand binding constants of nitrophorin 2 and its NO, histamine, and imidazole complexes.
Berry, R.E., Shokhirev, M.N., Ho, A.Y., Yang, F., Shokhireva, T.K., Zhang, H., Weichsel, A., Montfort, W.R., Walker, F.A.(2009) J Am Chem Soc 131: 2313-2327
- PubMed: 19175316
- DOI: https://doi.org/10.1021/ja808105d
- Primary Citation of Related Structures:
3FLL - PubMed Abstract:
Nitrophorins (NPs) are a group of NO-carrying heme proteins found in the saliva of a blood-sucking insect from tropical Central and South America, Rhodnius prolixus, the "kissing bug". NO is kept stable for long periods of time by binding it as an axial ligand to a ferriheme center. The fact that the nitrophorins are stabilized as Fe(III)-NO proteins is a unique property because most heme proteins are readily autoreduced by excess NO and bind NO to the Fe(II) heme irreversibly (K(d)s in the picomolar range). In contrast, the nitrophorins, as Fe(III) heme centers, have K(d)s in the micromolar to nanomolar range and thus allow NO to dissociate upon dilution following injection into the tissues of the victim. This NO can cause vasodilation and thereby allow more blood to be transported to the site of the wound. We prepared 13 site-directed mutants of three major nitrophorins, NP2, NP1, and NP4, to investigate the stabilization of the ferric-NO heme center and preservation of reversible binding that facilitates these proteins' NO storage, transport, and release functions. Of the mutations in which Glu and/or Asp were replaced by Ala, most of these carboxyls show a significant role stabilizing Fe(III)-NO over Fe(II)-NO, with buried E53 of NP2 or E55 of NP1 and NP4 being the most important and partially buried D29 of NP2 or D30 of NP4 being second in importance. The pK(a)s of the carboxyl groups studied vary significantly but all are largely deprotonated at pH 7.5 except E124.
Organizational Affiliation:
Department of Chemistry and Biochemistry, The University of Arizona, P.O. Box 210041, Tucson, Arizona 85721-0041, USA. berryr@email.arizona.edu