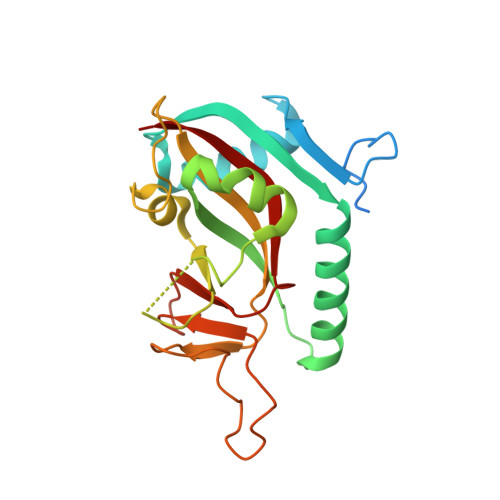
Structural Basis for Lack of ADP-ribosyltransferase Activity in Poly(ADP-ribose) Polymerase-13/Zinc Finger Antiviral Protein.
Karlberg, T., Klepsch, M., Thorsell, A.G., Andersson, C.D., Linusson, A., Schuler, H.(2015) J Biol Chem 290: 7336-7344
- PubMed: 25635049
- DOI: https://doi.org/10.1074/jbc.M114.630160
- Primary Citation of Related Structures:
2PQF, 2X5Y, 3BLJ, 3GEY, 4X52 - PubMed Abstract:
The mammalian poly(ADP-ribose) polymerase (PARP) family includes ADP-ribosyltransferases with diphtheria toxin homology (ARTD). Most members have mono-ADP-ribosyltransferase activity. PARP13/ARTD13, also called zinc finger antiviral protein, has roles in viral immunity and microRNA-mediated stress responses. PARP13 features a divergent PARP homology domain missing a PARP consensus sequence motif; the domain has enigmatic functions and apparently lacks catalytic activity. We used x-ray crystallography, molecular dynamics simulations, and biochemical analyses to investigate the structural requirements for ADP-ribosyltransferase activity in human PARP13 and two of its functional partners in stress granules: PARP12/ARTD12, and PARP15/BAL3/ARTD7. The crystal structure of the PARP homology domain of PARP13 shows obstruction of the canonical active site, precluding NAD(+) binding. Molecular dynamics simulations indicate that this closed cleft conformation is maintained in solution. Introducing consensus side chains in PARP13 did not result in 3-aminobenzamide binding, but in further closure of the site. Three-dimensional alignment of the PARP homology domains of PARP13, PARP12, and PARP15 illustrates placement of PARP13 residues that deviate from the PARP family consensus. Introducing either one of two of these side chains into the corresponding positions in PARP15 abolished PARP15 ADP-ribosyltransferase activity. Taken together, our results show that PARP13 lacks the structural requirements for ADP-ribosyltransferase activity.
Organizational Affiliation:
From the Structural Genomics Consortium and the Department of Medical Biochemistry and Biophysics, Karolinska Institutet, 17177 Stockholm, Sweden and.