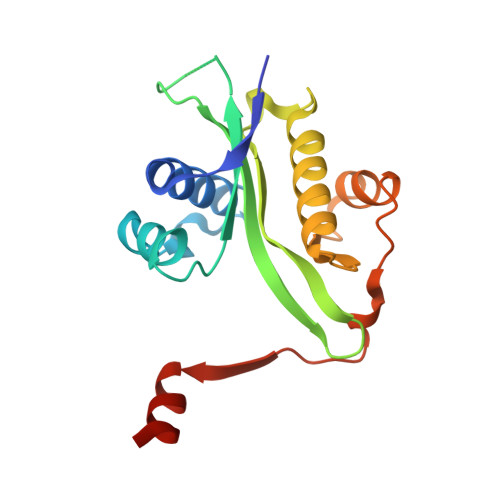
The crystal structure of spermidine/spermine N1-acetyltransferase in complex with spermine provides insights into substrate binding and catalysis.
Montemayor, E.J., Hoffman, D.W.(2008) Biochemistry 47: 9145-9153
- PubMed: 18690703
- DOI: https://doi.org/10.1021/bi8009357
- Primary Citation of Related Structures:
3BJ7, 3BJ8 - PubMed Abstract:
The enzyme spermidine/spermine N (1)-acetyltransferase (SSAT) catalyzes the transfer of acetyl groups from acetylcoenzyme A to spermidine and spermine, as part of a polyamine degradation pathway. This work describes the crystal structure of SSAT in complex with coenzyme A, with and without bound spermine. The complex with spermine provides a direct view of substrate binding by an SSAT and demonstrates structural plasticity near the active site of the enzyme. Associated water molecules bridge several of the intermolecular contacts between spermine and the enzyme and form a "proton wire" between the side chain of Glu92 and the N1 amine of spermine. A single water molecule can also be seen forming hydrogen bonds with the side chains of Glu92, Asp93, and the N4 amine of spermine. Site-directed mutation of Glu92 to glutamine had a detrimental effect on both substrate binding and catalysis and shifted the optimal pH for enzyme activity further into alkaline solution conditions, while mutation of Asp93 to asparagine affected both substrate binding and catalysis without changing the pH dependence of the enzyme. Considered together, the structural and kinetic data suggest that Glu92 functions as a catalytic base to drive an otherwise unfavorable deprotonation step at physiological pH.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Institute for Cellular and Molecular Biology, University of Texas, Austin, Texas 78712, USA.