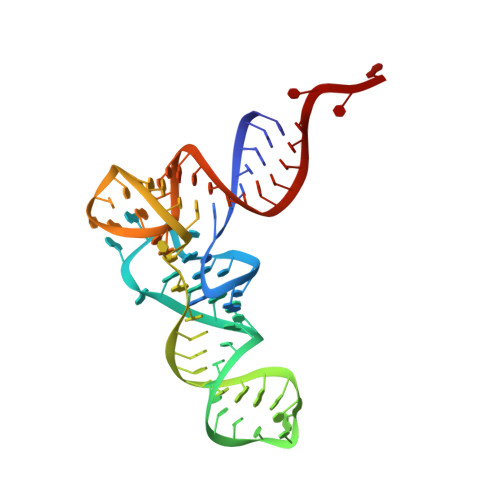
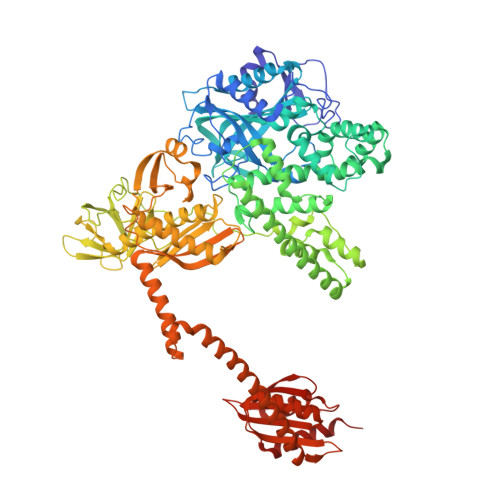
The selective tRNA aminoacylation mechanism based on a single G.U pair
Naganuma, M., Sekine, S., Chong, Y.E., Guo, M., Yang, X.L., Gamper, H., Hou, Y.M., Schimmel, P., Yokoyama, S.(2014) Nature 510: 507-511
- PubMed: 24919148
- DOI: https://doi.org/10.1038/nature13440
- Primary Citation of Related Structures:
3WQY, 3WQZ - PubMed Abstract:
Ligation of tRNAs with their cognate amino acids, by aminoacyl-tRNA synthetases, establishes the genetic code. Throughout evolution, tRNA(Ala) selection by alanyl-tRNA synthetase (AlaRS) has depended predominantly on a single wobble base pair in the acceptor stem, G3•U70, mainly on the kcat level. Here we report the crystal structures of an archaeal AlaRS in complex with tRNA(Ala) with G3•U70 and its A3•U70 variant. AlaRS interacts with both the minor- and the major-groove sides of G3•U70, widening the major groove. The geometry difference between G3•U70 and A3•U70 is transmitted along the acceptor stem to the 3'-CCA region. Thus, the 3'-CCA region of tRNA(Ala) with G3•U70 is oriented to the reactive route that reaches the active site, whereas that of the A3•U70 variant is folded back into the non-reactive route. This novel mechanism enables the single wobble pair to dominantly determine the specificity of tRNA selection, by an approximate 100-fold difference in kcat.
Organizational Affiliation:
1] RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan [2] Department of Biophysics and Biochemistry and Laboratory of Structural Biology, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan [3] RIKEN Structural Biology Laboratory, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan.