Crystal structure of GTPase domain fused with minimal stalks from human dynamin-1-like protein (Dlp1) in complex with several nucleotide analogues
Kishida, H., Sugio, S.(2013) Curr Top Pept Protein Res 14: 67-77
Experimental Data Snapshot
(2013) Curr Top Pept Protein Res 14: 67-77
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Dynamin-1-like protein | 364 | Homo sapiens | Mutation(s): 0 Gene Names: DNM1L, DLP1, DRP1 EC: 3.6.5.5 | 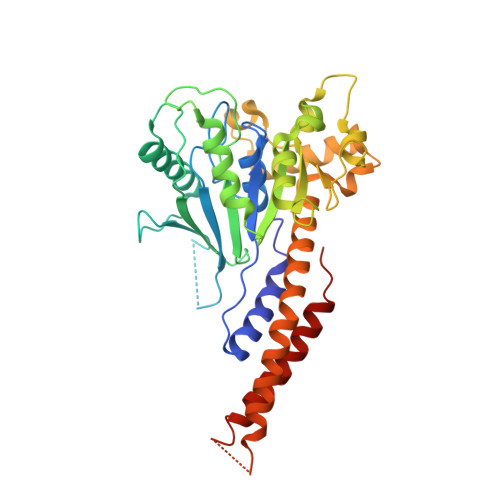 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for O00429 (Homo sapiens) Explore O00429 Go to UniProtKB: O00429 | |||||
PHAROS: O00429 GTEx: ENSG00000087470 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O00429 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GCP Query on GCP | C [auth A], E [auth B] | PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER C11 H18 N5 O13 P3 PHBDHXOBFUBCJD-KQYNXXCUSA-N |  | ||
| PG4 Query on PG4 | G [auth B], H [auth B] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| CA Query on CA | I [auth B] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| MG Query on MG | D [auth A], F [auth B] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.24 | α = 90 |
| b = 109.07 | β = 90 |
| c = 128.3 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHASER | phasing |
| REFMAC | refinement |
| XDS | data reduction |
| XSCALE | data scaling |