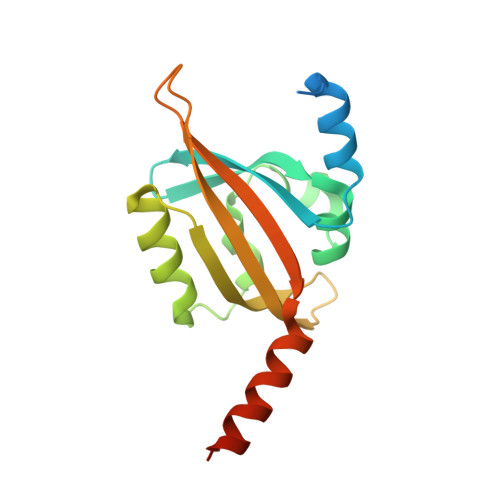
Structural Basis for the Slow Dark Recovery of a Full-Length LOV Protein from Pseudomonas putida.
Circolone, F., Granzin, J., Jentzsch, K., Drepper, T., Jaeger, K.E., Willbold, D., Krauss, U., Batra-Safferling, R.(2012) J Mol Biol 417: 362-374
- PubMed: 22326872
- DOI: https://doi.org/10.1016/j.jmb.2012.01.056
- Primary Citation of Related Structures:
3SW1 - PubMed Abstract:
Blue-light photoreceptors containing light–oxygen–voltage (LOV) domains regulate a myriad of different physiological responses in both eukaryotes and prokaryotes. Their light sensitivity is intricately linked to the photochemistry of the non-covalently bound flavin mononucleotide (FMN) chromophore that forms a covalent adduct with a conserved cysteine residue in the LOV domain upon illumination with blue light. All LOV domains undergo the same primary photochemistry leading to adduct formation; however, considerable variation is found in the lifetime of the adduct state that varies from seconds to several hours. The molecular mechanism underlying this variation among the structurally conserved LOV protein family is not well understood. Here, we describe the structural characterization of PpSB1-LOV, a very slow cycling full-length LOV protein from the Gram-negative bacterium Pseudomonas putida KT2440. Its crystal structure reveals a novel dimer interface that is mediated by N- and C-terminal auxiliary structural elements and a unique cluster of four arginine residues coordinating with the FMN-phosphate moiety. Site-directed mutagenesis of two arginines (R61 and R66) in PpSB1-LOV resulted in acceleration of the dark recovery reaction approximately by a factor of 280. The presented structural and biochemical data suggest a direct link between structural features and the slow dark recovery observed for PpSB1-LOV. The overall structural arrangement of PpSB1-LOV, together with a complementary phylogenetic analysis, highlights a common ancestry of bacterial LOV photoreceptors and Per-ARNT-Sim chemosensors.
Organizational Affiliation:
Institut für Molekulare Enzymtechnologie, Heinrich-Heine-Universität Düsseldorf, D-52426 Jülich, Germany.