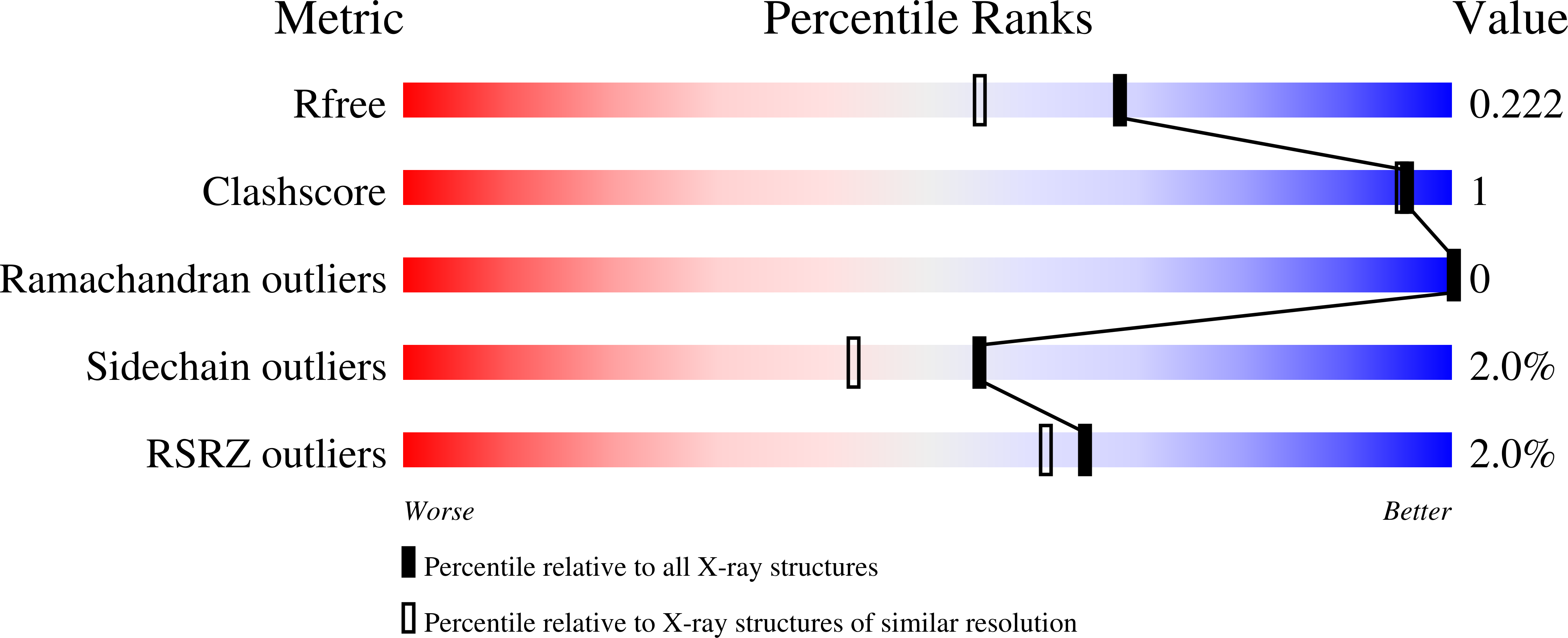
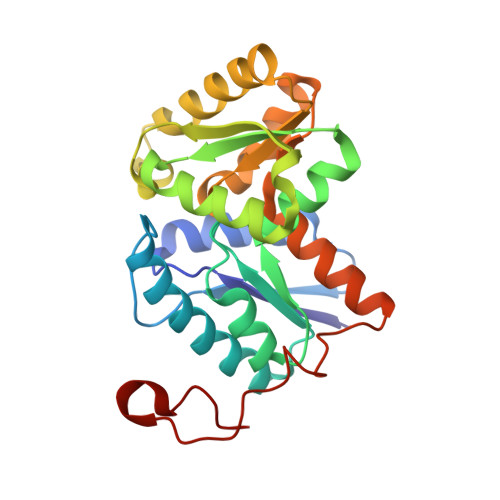
Characterization of the Structure and Function of Klebsiella pneumoniae Allantoin Racemase.
French, J.B., Neau, D.B., Ealick, S.E.(2011) J Mol Biol 410: 447-460
- PubMed: 21616082
- DOI: https://doi.org/10.1016/j.jmb.2011.05.016
- Primary Citation of Related Structures:
3QVJ, 3QVK, 3QVL - PubMed Abstract:
The oxidative catabolism of uric acid produces 5-hydroxyisourate (HIU), which is further degraded to (S)-allantoin by two enzymes, HIU hydrolase and 2-oxo-4-hydroxy-4-carboxy-5-ureidoimidazoline decarboxylase. The intermediates of the latter two reactions, HIU and 2-oxo-4-hydroxy-4-carboxy-5-ureidoimidazoline, are unstable in solution and decay nonstereospecifically to allantoin. In addition, nonenzymatic racemization of allantoin has been shown to occur at physiological pH. Since the further breakdown of allantoin is catalyzed by allantoinase, an enzyme that is specific for (S)-allantoin, an allantoin racemase is necessary for complete and efficient catabolism of uric acid. In this work, we characterize the structure and activity of allantoin racemase from Klebsiella pneumoniae (KpHpxA). In addition to an unliganded structure solved using selenomethionyl single-wavelength anomalous dispersion, structures of C79S/C184S KpHpxA in complex with allantoin and with 5-acetylhydantoin are presented. These structures reveal several important features of the active site including an oxyanion hole and a polar binding pocket that interacts with the ureido tail of allantoin and serves to control the orientation of the hydantoin ring. The ability of KpHpxA to interconvert the (R)- and (S)-enantiomers of allantoin is demonstrated, and analysis of the steady-state kinetics of KpHpxA yielded a k(cat)/K(m) of 6.0 × 10(5) M(-1) s(-1). Mutation of either of the active-site cysteines, Cys79 or Cys184, to serine inactivates this enzyme. The data presented provide new insights into the activity and substrate specificity of this enzyme and enable us to propose a mechanism for catalysis that is consistent with the two-base mechanism observed in other members of the aspartate/glutamate family.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Cornell University, Ithaca, NY 14853, USA.