CRYSTAL STRUCTURE OF D-MANNONATE DEHYDRATASE FROM CHROMOHALOBACTER SALEXIGENS complexed with MG,D-Mannonate and 2-keto-3-deoxy-D-Gluconate
Fedorov, A.A., Fedorov, E.V., Wichelecki, D., Gerlt, J.A., Almo, S.C.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mandelate racemase/muconate lactonizing enzyme | 405 | Chromohalobacter salexigens | Mutation(s): 0 Gene Names: Csal_2974 | 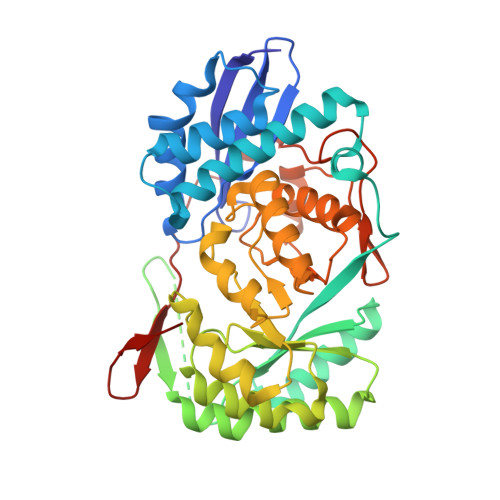 | |
UniProt | |||||
Find proteins for Q1QT89 (Chromohalobacter salexigens (strain ATCC BAA-138 / DSM 3043 / CIP 106854 / NCIMB 13768 / 1H11)) Explore Q1QT89 Go to UniProtKB: Q1QT89 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q1QT89 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CS2 Query on CS2 | J [auth A], L [auth B], R [auth E], T [auth F] | D-MANNONIC ACID C6 H12 O7 RGHNJXZEOKUKBD-MBMOQRBOSA-N |  | ||
| KDG Query on KDG | N [auth C], P [auth D], V [auth G], X [auth H] | 2-KETO-3-DEOXYGLUCONATE C6 H10 O6 WPAMZTWLKIDIOP-WVZVXSGGSA-N |  | ||
| MG Query on MG | I [auth A] K [auth B] M [auth C] O [auth D] Q [auth E] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 110.242 | α = 90 |
| b = 167.263 | β = 90 |
| c = 168.863 | γ = 90 |
| Software Name | Purpose |
|---|---|
| ADSC | data collection |
| BALBES | phasing |
| PHENIX | refinement |
| DENZO | data reduction |
| SCALEPACK | data scaling |