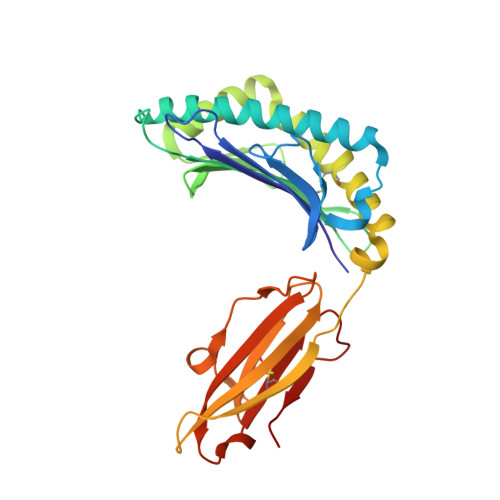
Identification and structural definition of H5-specific CTL epitopes restricted by HLA-A*0201 derived from the H5N1 subtype of influenza A viruses
Sun, Y., Liu, J., Yang, M., Gao, F., Zhou, J., Kitamura, Y., Gao, B., Tien, P., Shu, Y., Iwamoto, A., Chen, Z., Gao, G.F.(2010) J Gen Virol 91: 919-930
- PubMed: 19955560
- DOI: https://doi.org/10.1099/vir.0.016766-0
- Primary Citation of Related Structures:
3MGO, 3MGT - PubMed Abstract:
The haemagglutinin (HA) glycoprotein of influenza A virus is a major antigen that initiates humoral immunity against infection; however, the cellular immune response against HA is poorly understood. Furthermore, HA-derived cytotoxic T-lymphocyte (CTL) epitopes are relatively rare in comparison to other internal gene products. Here, CTL epitopes of the HA serotype H5 protein were screened. By using in silico prediction, in vitro refolding and a T2 cell-binding assay, followed by immunization of HLA-A2.1/K(b) transgenic mice, an HLA-A*0201-restricted decameric epitope, RI-10 (H5 HA205-214, RLYQNPTTYI), was shown to elicit a robust CTL epitope-specific response. In addition, RI-10 and its variant, KI-10 (KLYQNPTTYI), were also demonstrated to be able to induce a higher CTL epitope-specific response than the influenza A virus dominant CTL epitope GL-9 (GILGFVFTL) in peripheral blood mononuclear cells of HLA-A*0201-positive patients who had recovered from H5N1 virus infection. Furthermore, the crystal structures of RI-10-HLA-A*0201 and KI-10-HLA-A*0201 complexes were determined at 2.3 and 2.2 A resolution, respectively, showing typical HLA-A*0201-restricted epitopes. The conformations of RI-10 and KI-10 in the antigen-presenting grooves in crystal structures of the two complexes show significant differences, despite their nearly identical sequences. These results provide implications for the discovery of diagnostic markers and the design of novel influenza vaccines.
Organizational Affiliation:
CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences (CAS), Beijing, PR China.