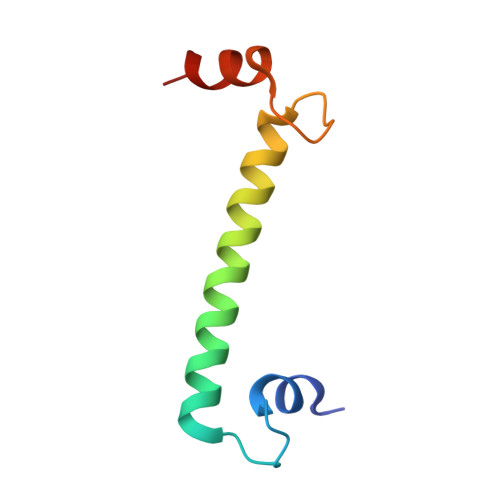
Crystal structure and trimer-monomer transition of N-terminal domain of EhCaBP1 from Entamoeba histolytica
Kumar, S., Ahmad, E., Mansuri, M.S., Kumar, S., Jain, R., Khan, R.H., Gourinath, S.(2010) Biophys J 98: 2933-2942
- PubMed: 20550906
- DOI: https://doi.org/10.1016/j.bpj.2010.03.048
- Primary Citation of Related Structures:
3LI6 - PubMed Abstract:
EhCaBP1 is a well-characterized calcium binding protein from Entamoeba histolytica with four canonical EF-hand motifs. The crystal structure of EhCaBP1 reveals the trimeric organization of N-terminal domain. The solution structure obtained at pH 6.0 indicated its monomeric nature, similar to that of calmodulin. Recent domain-wise studies showed clearly that the N-terminal domain of EhCaBP1 is capable of performing most of the functions of the full-length protein. Additionally, the mode of target binding in the trimer is similar to that found in calmodulin. To study the dynamic nature of this protein and further validate the trimerization of N-terminal domain at physiological conditions, the crystal structure of N-terminal domain was determined at 2.5 A resolution. The final structure consists of EF-1 and EF-2 motifs separated by a long straight helix as seen in the full-length protein. The spectroscopic and stability studies, like far and near-ultraviolet circular dichroism spectra, intrinsic and extrinsic fluorescence spectra, acrylamide quenching, thermal denaturation, and dynamic light scattering, provided clear evidence for a conversion from trimeric state to monomeric state. As the pH was lowered from the physiological pH, a dynamic trimer-monomer transition was observed. The trimeric state and monomeric state observed in spectroscopic studies may represent the x-ray and NMR structures of the EhCaBP1. At pH 6.0, the endogenous kinase activation function was almost lost, indicating that the monomeric state of the protein, where EF-hand motifs are far apart, is not a functional state.
Organizational Affiliation:
School of Life Sciences, Jawaharlal Nehru University, New Delhi, India.