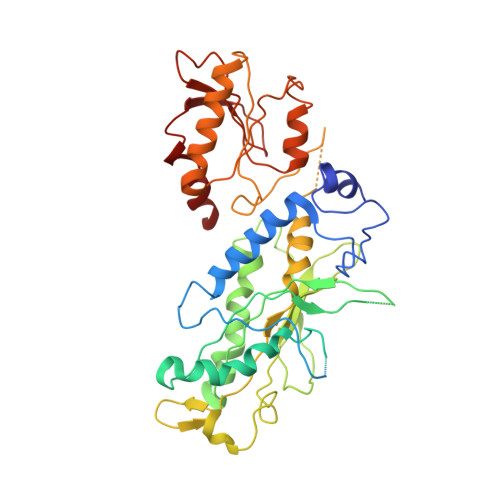
Structural insight into processive human mitochondrial DNA synthesis and disease-related polymerase mutations.
Lee, Y.S., Kennedy, W.D., Yin, Y.W.(2009) Cell 139: 312-324
- PubMed: 19837034
- DOI: https://doi.org/10.1016/j.cell.2009.07.050
- Primary Citation of Related Structures:
3IKL, 3IKM - PubMed Abstract:
Human mitochondrial DNA polymerase (Pol gamma) is the sole replicase in mitochondria. Pol gamma is vulnerable to nonselective antiretroviral drugs and is increasingly associated with mutations found in patients with mitochondriopathies. We determined crystal structures of the human heterotrimeric Pol gamma holoenzyme and, separately, a variant of its processivity factor, Pol gammaB. The holoenzyme structure reveals an unexpected assembly of the mitochondrial DNA replicase where the catalytic subunit Pol gammaA interacts with its processivity factor primarily via a domain that is absent in all other DNA polymerases. This domain provides a structural module for supporting both the intrinsic processivity of the catalytic subunit alone and the enhanced processivity of holoenzyme. The Pol gamma structure also provides a context for interpreting the phenotypes of disease-related mutations in the polymerase and establishes a foundation for understanding the molecular basis of toxicity of anti-retroviral drugs targeting HIV reverse transcriptase.
Organizational Affiliation:
Institute for Cellular and Molecular Biology, University of Texas at Austin, Austin, TX 78712, USA.