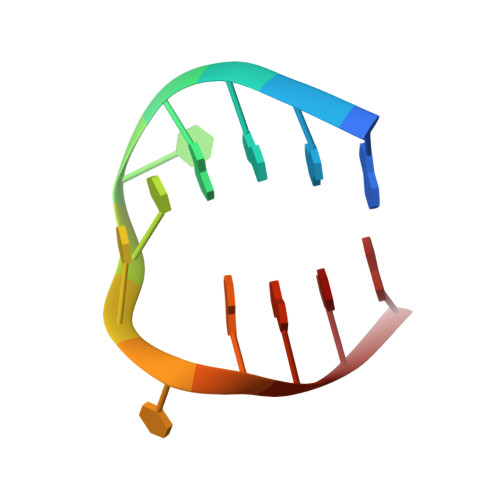
Selectivity in Ligand Recognition of G-Quadruplex Loops.
Campbell, N.H., Patel, M., Tofa, A.B., Ghosh, R., Parkinson, G.N., Neidle, S.(2009) Biochemistry 48: 1675-1680
- PubMed: 19173611
- DOI: https://doi.org/10.1021/bi802233v
- Primary Citation of Related Structures:
3EM2, 3EQW, 3ERU, 3ES0, 3ET8, 3EUI, 3EUM - PubMed Abstract:
A series of disubstituted acridine ligands have been cocrystallized with a bimolecular DNA G-quadruplex. The ligands have a range of cyclic amino end groups of varying size. The crystal structures show that the diagonal loop in this quadruplex results in a large cavity for these groups, in contrast to the steric constraints imposed by propeller loops in human telomeric quadruplexes. We conclude that the nature of the loop has a significant influence on ligand selectivity for particular quadruplex folds.
Organizational Affiliation:
Cancer Research UK Biomolecular Structure Group, The School of Pharmacy, University of London, 29-39 Brunswick Square, London WC1N 1AX, UK.