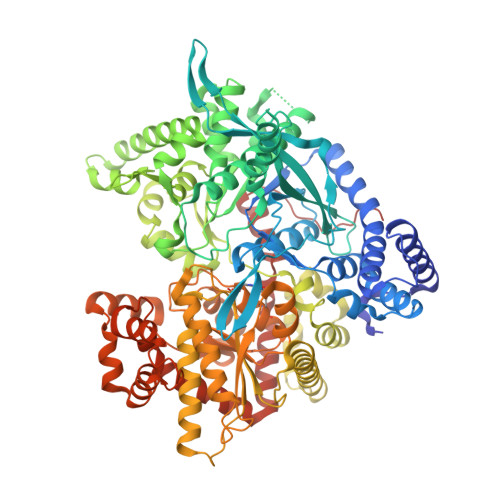
Tracking structure activity relationships of glycogen phosphorylase inhibitors: synthesis, kinetic and crystallographic evaluation of analogues of N-(-D-glucopyranosyl)-N'-oxamides
Czifrak, K., Kyritsi, C., Felfoldi, N., Dosca, T., Gergely, P., Chrysina, E.D., Siafaka-Kapadai, A., Leonidas, D.D., Zographos, S.E., Oikonomakos, N.G.To be published.