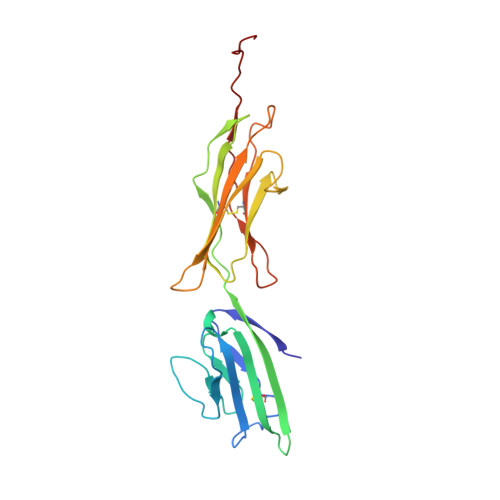
Structural basis for ligand recognition and activation of RAGE.
Koch, M., Chitayat, S., Dattilo, B.M., Schiefner, A., Diez, J., Chazin, W.J., Fritz, G.(2010) Structure 18: 1342-1352
- PubMed: 20947022
- DOI: https://doi.org/10.1016/j.str.2010.05.017
- Primary Citation of Related Structures:
3CJJ - PubMed Abstract:
The receptor for advanced glycation end products (RAGE) is a pattern recognition receptor involved in inflammatory processes and is associated with diabetic complications, tumor outgrowth, and neurodegenerative disorders. RAGE induces cellular signaling events upon binding of a variety of ligands, such as glycated proteins, amyloid-β, HMGB1, and S100 proteins. The X-ray crystal structure of the VC1 ligand-binding region of the human RAGE ectodomain was determined at 1.85 Å resolution. The VC1 ligand-binding surface was mapped onto the structure from titrations with S100B monitored by heteronuclear NMR spectroscopy. These NMR chemical shift perturbations were used as input for restrained docking calculations to generate a model for the VC1-S100B complex. Together, the arrangement of VC1 molecules in the crystal and complementary biochemical studies suggest a role for self-association in RAGE function. Our results enhance understanding of the functional outcomes of S100 protein binding to RAGE and provide insight into mechanistic models for how the receptor is activated.
Organizational Affiliation:
Department of Biology, University of Konstanz, 78457 Konstanz, Germany.