Structural basis for RNA-silencing suppression by Tomato aspermy virus protein 2b
Chen, H.-Y., Yang, J., Lin, C., Yuan, Y.A.(2008) EMBO Rep 9: 754-760
- PubMed: 18600235
- DOI: https://doi.org/10.1038/embor.2008.118
- Primary Citation of Related Structures:
2ZI0 - PubMed Abstract:
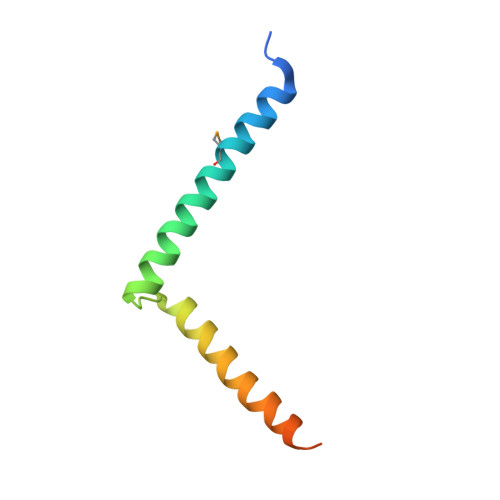
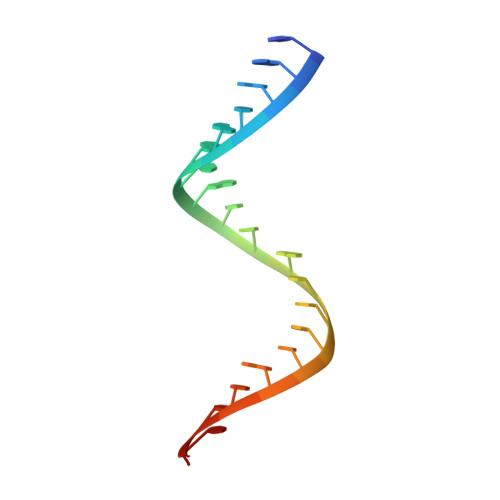
The 2b proteins encoded by cucumovirus act as post-transcriptional gene silencing suppressors to counter host defence during infection. Here we report the crystal structure of Tomato aspermy virus 2b (TAV2b) protein bound to a 19 bp small interfering RNA (siRNA) duplex. TAV2b adopts an all alpha-helix structure and forms a homodimer to measure siRNA duplex in a length-preference mode. TAV2b has a pair of hook-like structures to recognize simultaneously two alpha-helical turns of A-form RNA duplex by fitting its alpha-helix backbone into two adjacent major grooves of siRNA duplex. The conserved pi-stackings between tryptophan and the 5'-terminal base of siRNA duplex from both ends enhance the recognition. TAV2b further oligomerizes to form a dimer of dimers through the conserved leucine-zipper-like motif at its amino-terminal alpha-helix. Biochemical experiments suggest that TAV2b might interfere with the post-transcriptional gene silencing pathway by directly binding to siRNA duplex.
Organizational Affiliation:
Host-Pathogen Interaction Group, National University of Singapore, Singapore.