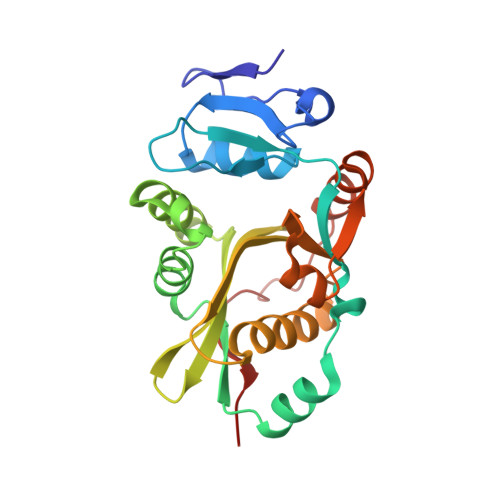
Protein-based peptide-bond formation by aminoacyl-tRNA protein transferase
Watanabe, K., Toh, Y., Suto, K., Shimizu, Y., Oka, N., Wada, T., Tomita, K.(2007) Nature 449: 867-871
- PubMed: 17891155
- DOI: https://doi.org/10.1038/nature06167
- Primary Citation of Related Structures:
2Z3K, 2Z3L, 2Z3M, 2Z3N, 2Z3O, 2Z3P - PubMed Abstract:
Eubacterial leucyl/phenylalanyl-tRNA protein transferase (LF-transferase) catalyses peptide-bond formation by using Leu-tRNA(Leu) (or Phe-tRNA(Phe)) and an amino-terminal Arg (or Lys) of a protein, as donor and acceptor substrates, respectively. However, the catalytic mechanism of peptide-bond formation by LF-transferase remained obscure. Here we determine the structures of complexes of LF-transferase and phenylalanyl adenosine, with and without a short peptide bearing an N-terminal Arg. Combining the two separate structures into one structure as well as mutation studies reveal the mechanism for peptide-bond formation by LF-transferase. The electron relay from Asp 186 to Gln 188 helps Gln 188 to attract a proton from the alpha-amino group of the N-terminal Arg of the acceptor peptide. This generates the attacking nucleophile for the carbonyl carbon of the aminoacyl bond of the aminoacyl-tRNA, thus facilitating peptide-bond formation. The protein-based mechanism for peptide-bond formation by LF-transferase is similar to the reverse reaction of the acylation step observed in the peptide hydrolysis reaction by serine proteases.
Organizational Affiliation:
Institute of Biological Resources and Functions, National Institute of Advanced Industrial Sciences and Technology, 1-1-1, Higashi, Tsukuba-shi, Ibaraki 305-8566, Japan.