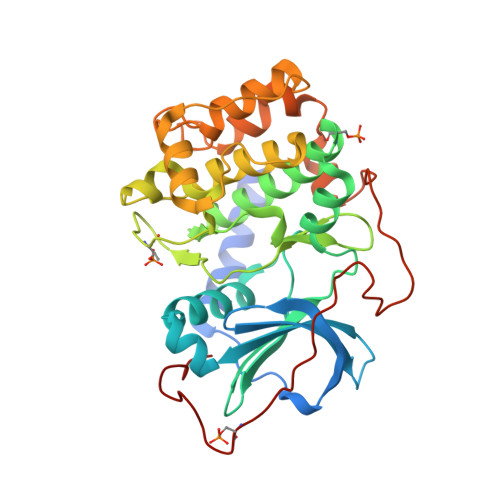
Identification of Inhibitors of Protein Kinase B Using Fragment-Based Lead Discovery
Saxty, G., Woodhead, S.J., Berdini, V., Davies, T.G., Verdonk, M.L., Wyatt, P.G., Boyle, R.G., Barford, D., Downham, R., Garrett, M.D., Carr, R.A.(2007) J Med Chem 50: 2293
- PubMed: 17451234
- DOI: https://doi.org/10.1021/jm070091b
- Primary Citation of Related Structures:
2UW3, 2UW4, 2UW5, 2UW6, 2UW7, 2UW8, 2UW9 - PubMed Abstract:
Using fragment-based screening techniques, 5-methyl-4-phenyl-1H-pyrazole (IC50 80 microM) was identified as a novel, low molecular weight inhibitor of protein kinase B (PKB). Herein we describe the rapid elaboration of highly potent and ligand efficient analogues using a fragment growing approach. Iterative structure-based design was supported by protein-ligand structure determinations using a PKA-PKB "chimera" and a final protein-ligand structure of a lead compound in PKBbeta itself.