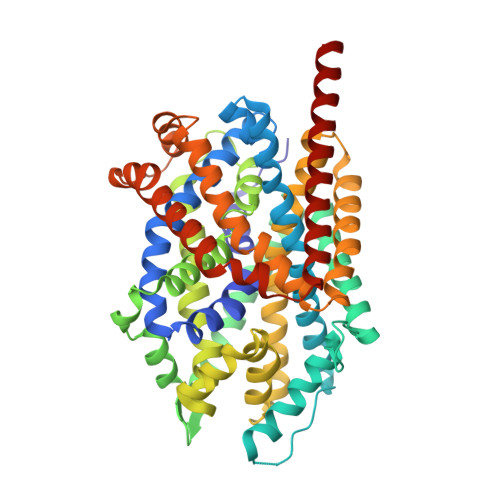
LeuT-desipramine structure reveals how antidepressants block neurotransmitter reuptake.
Zhou, Z., Zhen, J., Karpowich, N.K., Goetz, R.M., Law, C.J., Reith, M.E., Wang, D.N.(2007) Science 317: 1390-1393
- PubMed: 17690258
- DOI: https://doi.org/10.1126/science.1147614
- Primary Citation of Related Structures:
2QJU - PubMed Abstract:
Tricyclic antidepressants exert their pharmacological effect-inhibiting the reuptake of serotonin, norepinephrine, and dopamine-by directly blocking neurotransmitter transporters (SERT, NET, and DAT, respectively) in the presynaptic membrane. The drug-binding site and the mechanism of this inhibition are poorly understood. We determined the crystal structure at 2.9 angstroms of the bacterial leucine transporter (LeuT), a homolog of SERT, NET, and DAT, in complex with leucine and the antidepressant desipramine. Desipramine binds at the inner end of the extracellular cavity of the transporter and is held in place by a hairpin loop and by a salt bridge. This binding site is separated from the leucine-binding site by the extracellular gate of the transporter. By directly locking the gate, desipramine prevents conformational changes and blocks substrate transport. Mutagenesis experiments on human SERT and DAT indicate that both the desipramine-binding site and its inhibition mechanism are probably conserved in the human neurotransmitter transporters.
Organizational Affiliation:
Kimmel Center for Biology and Medicine at the Skirball Institute of Biomolecular Medicine and Department of Cell Biology, New York University School of Medicine, 540 First Avenue, New York, NY 10016, USA.