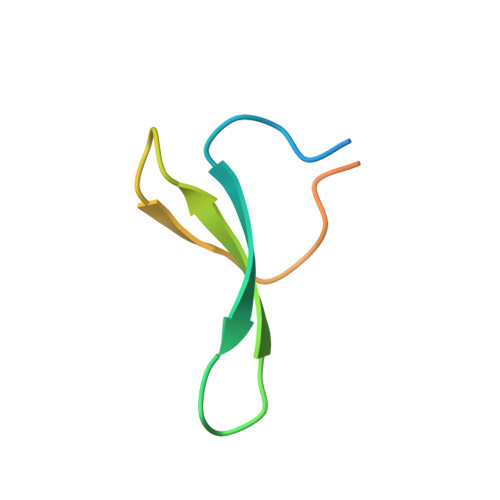
Structural Basis of the Activation and Degradation Mechanisms of the E3 Ubiquitin Ligase Nedd4L.
Escobedo, A., Gomes, T., Aragon, E., Martin-Malpartida, P., Ruiz, L., Macias, M.J.(2014) Structure 22: 1446-1457
- PubMed: 25295397
- DOI: https://doi.org/10.1016/j.str.2014.08.016
- Primary Citation of Related Structures:
2MPT - PubMed Abstract:
We investigated the mechanisms of activation and degradation of the E3 ubiquitin ligase Nedd4L combining the available biochemical information with complementary biophysical techniques. Using nuclear magnetic resonance spectroscopy, we identified that the C2 domain binds Ca(2+) and inositol 1,4,5-trisphosphate (IP3) using the same interface that is used to interact with the HECT domain. Thus, we propose that the transition from the closed to the active form is regulated by a competition of IP3 and Ca(2+) with the HECT domain for binding to the C2 domain. We performed relaxation experiments and molecular dynamic simulations to determine the flexibility of the HECT structure and observed that its conserved PY motif can become solvent-exposed when the unfolding process is initiated. The structure of the WW3 domain bound to the HECT-PY site reveals the details of this interaction, suggesting a possible auto-ubquitination mechanism using two molecules, a partially unfolded one and a fully functional Nedd4L counterpart.
Organizational Affiliation:
Institute for Research in Biomedicine (IRB Barcelona), Baldiri Reixac 10, 08028 Barcelona, Spain.