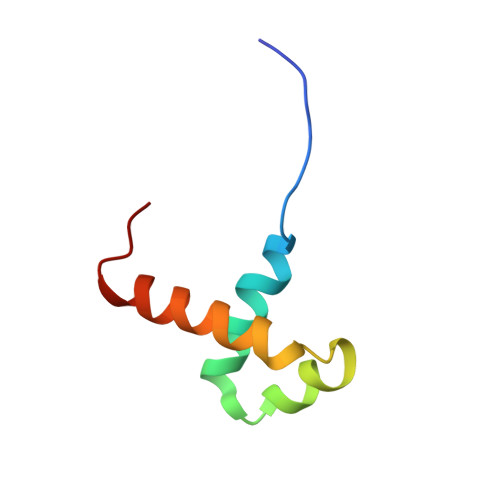
NMR structural analysis of Sleeping Beauty transposase binding to DNA.
Carpentier, C.E., Schreifels, J.M., Aronovich, E.L., Carlson, D.F., Hackett, P.B., Nesmelova, I.V.(2014) Protein Sci 23: 23-33
- PubMed: 24243759
- DOI: https://doi.org/10.1002/pro.2386
- Primary Citation of Related Structures:
2M8E - PubMed Abstract:
The Sleeping Beauty (SB) transposon is the most widely used DNA transposon in genetic applications and is the only DNA transposon thus far in clinical trials for human gene therapy. In the absence of atomic level structural information, the development of SB transposon relied primarily on the biochemical and genetic homology data. While these studies were successful and have yielded hyperactive transposases, structural information is needed to gain a mechanistic understanding of transposase activity and guides to further improvement. We have initiated a structural study of SB transposase using Nuclear Magnetic Resonance (NMR) and Circular Dichroism (CD) spectroscopy to investigate the properties of the DNA-binding domain of SB transposase in solution. We show that at physiologic salt concentrations, the SB DNA-binding domain remains mostly unstructured but its N-terminal PAI subdomain forms a compact, three-helical structure with a helix-turn-helix motif at higher concentrations of NaCl. Furthermore, we show that the full-length SB DNA-binding domain associates differently with inner and outer binding sites of the transposon DNA. We also show that the PAI subdomain of SB DNA-binding domain has a dominant role in transposase's attachment to the inverted terminal repeats of the transposon DNA. Overall, our data validate several earlier predictions and provide new insights on how SB transposase recognizes transposon DNA.