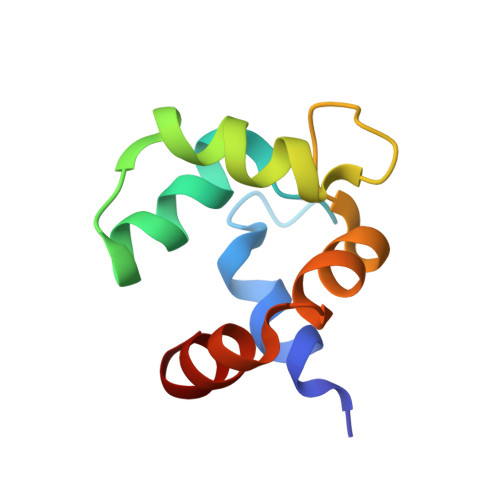
Solution structures of polcalcin Phl p 7 in three ligation states: Apo-, hemi-Mg(2+) -bound, and fully Ca(2+) -bound.
Henzl, M.T., Sirianni, A.G., Wycoff, W.G., Tan, A., Tanner, J.J.(2013) Proteins 81: 300-315
- PubMed: 23011803
- DOI: https://doi.org/10.1002/prot.24186
- Primary Citation of Related Structures:
2LVI, 2LVJ, 2LVK - PubMed Abstract:
Polcalcins are small EF-hand proteins believed to assist in regulating pollen-tube growth. Phl p 7, from timothy grass (Phleum pratense), crystallizes as a domain-swapped dimer at low pH. This study describes the solution structures of the recombinant protein in buffered saline at pH 6.0, containing either 5.0 mM EDTA, 5.0 mM Mg(2+), or 100 μM Ca(2+). Phl p 7 is monomeric in all three ligation states. In the apo-form, both EF-hand motifs reside in the closed conformation, with roughly antiparallel N- and C-terminal helical segments. In 5.0 mM Mg(2+), the divalent ion is bound by EF-hand 2, perturbing interhelical angles and imposing more regular helical structure. The structure of Ca(2+)-bound Phl p 7 resembles that previously reported for Bet v 4-likewise exposing apolar surface to the solvent. Occluded in the apo- and Mg(2+)-bound forms, this surface presumably provides the docking site for Phl p 7 targets. Unlike Bet v 4, EF-hand 2 in Phl p 7 includes five potential anionic ligands, due to replacement of the consensus serine residue at -x (residue 55 in Phl p 7) with aspartate. In the Phl p 7 crystal structure, D55 functions as a helix cap for helix D. In solution, however, D55 apparently serves as a ligand to the bound Ca(2+). When Mg(2+) resides in site 2, the D55 carboxylate withdraws to a distance consistent with a role as an outer-sphere ligand. (15)N relaxation data, collected at 600 MHz, indicate that backbone mobility is limited in all three ligation states.
Organizational Affiliation:
Department of Biochemistry, University of Missouri, Columbia, Missouri 65211, USA. henzlm@missouri.edu