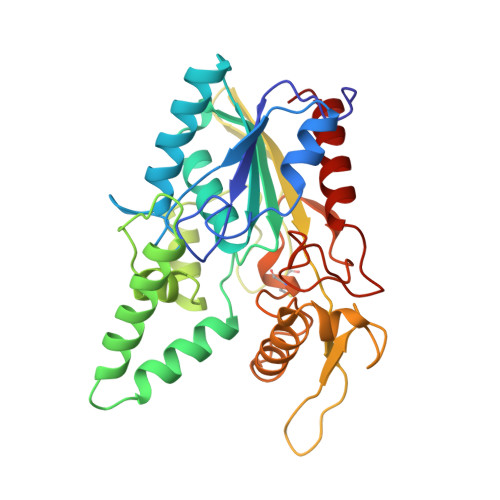
The open conformation of a Pseudomonas lipase.
Schrag, J.D., Li, Y., Cygler, M., Lang, D., Burgdorf, T., Hecht, H.J., Schmid, R., Schomburg, D., Rydel, T.J., Oliver, J.D., Strickland, L.C., Dunaway, C.M., Larson, S.B., Day, J., McPherson, A.(1997) Structure 5: 187-202
- PubMed: 9032074
- DOI: https://doi.org/10.1016/s0969-2126(97)00178-0
- Primary Citation of Related Structures:
2LIP, 3LIP - PubMed Abstract:
. The interfacial activation of lipases results primarily from conformational changes in the enzymes which expose the active site and provide a hydrophobic surface for interaction with the lipid substrate. Comparison of the crystallization conditions used and the structures observed for a variety of lipases suggests that the enzyme conformation is dependent on solution conditions. Pseudomonas cepacia lipase (PCL) was crystallized in conditions from which the open, active conformation of the enzyme was expected. Its three-dimensional structure was determined independently in three different laboratories and was compared with the previously reported closed conformations of the closely related lipases from Pseudomonas glumae (PGL) and Chromobacterium viscosum (CVL). These structures provide new insights into the function of this commercially important family of lipases.
Organizational Affiliation:
Biotechnology Research Institute, NRC of Canada, 6100 Royalmount Ave. Montreal, Quebec H4P 2R2, Canada. Joe.Schrag@nrc.ca