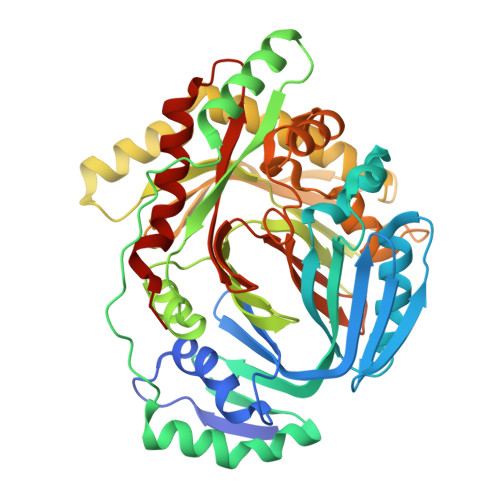
The Structure of MbtI from Mycobacterium tuberculosis, the First Enzyme in the Biosynthesis of the Siderophore Mycobactin, Reveals It To Be a Salicylate Synthase
Harrison, A.J., Gardenborg, T., Yu, M., Middleditch, M., Ramsay, R.J., Baker, E.N., Lott, J.S.(2006) J Bacteriol 188: 6081-6091
- PubMed: 16923875
- DOI: https://doi.org/10.1128/JB.00338-06
- Primary Citation of Related Structures:
2G5F - PubMed Abstract:
The ability to acquire iron from the extracellular environment is a key determinant of pathogenicity in mycobacteria. Mycobacterium tuberculosis acquires iron exclusively via the siderophore mycobactin T, the biosynthesis of which depends on the production of salicylate from chorismate. Salicylate production in other bacteria is either a two-step process involving an isochorismate synthase (chorismate isomerase) and a pyruvate lyase, as observed for Pseudomonas aeruginosa, or a single-step conversion catalyzed by a salicylate synthase, as with Yersinia enterocolitica. Here we present the structure of the enzyme MbtI (Rv2386c) from M. tuberculosis, solved by multiwavelength anomalous diffraction at a resolution of 1.8 A, and biochemical evidence that it is the salicylate synthase necessary for mycobactin biosynthesis. The enzyme is critically dependent on Mg2+ for activity and produces salicylate via an isochorismate intermediate. MbtI is structurally similar to salicylate synthase (Irp9) from Y. enterocolitica and the large subunit of anthranilate synthase (TrpE) and shares the overall architecture of other chorismate-utilizing enzymes, such as the related aminodeoxychorismate synthase PabB. Like Irp9, but unlike TrpE or PabB, MbtI is neither regulated by nor structurally stabilized by bound tryptophan. The structure of MbtI is the starting point for the design of inhibitors of siderophore biosynthesis, which may make useful lead compounds for the production of new antituberculosis drugs, given the strong dependence of pathogenesis on iron acquisition in M. tuberculosis.
Organizational Affiliation:
Centre for Molecular Biodiscovery, University of Auckland, Auckland, New Zealand.