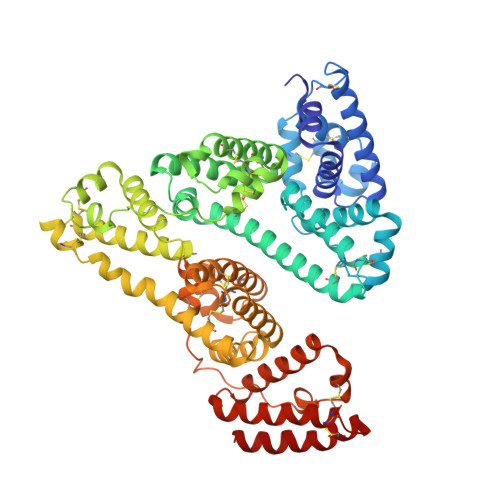
Structural Basis of Binding of Fluorescent, Site-Specific Dansylated Amino Acids to Human Serum Albumin.
Ryan, A.J., Ghuman, J., Zunszain, P.A., Chung, C.W., Curry, S.(2011) J Struct Biol 174: 84
- PubMed: 20940056
- DOI: https://doi.org/10.1016/j.jsb.2010.10.004
- Primary Citation of Related Structures:
2XSI, 2XVQ, 2XVU, 2XVV, 2XVW, 2XW0, 2XW1 - PubMed Abstract:
Human serum albumin (HSA) has two primary binding sites for drug molecules. These sites selectively bind different dansylated amino acid compounds, which-due to their intrinsic fluorescence-have long been used as specific markers for the drug pockets on HSA. We present here the co-crystal structures of HSA in complex with six dansylated amino acids that are specific for either drug site 1 (dansyl-l-asparagine, dansyl-l-arginine, dansyl-l-glutamate) or drug site 2 (dansyl-l-norvaline, dansyl-l-phenylalanine, dansyl-l-sarcosine). Our results explain the structural basis of the site-specificity of different dansylated amino acids. They also show that fatty acid binding has only a modest effect on binding of dansylated amino acids to drug site 1 and identify the location of secondary binding sites.
Organizational Affiliation:
Biophysics Section, Blackett Laboratory, Imperial College, Exhibition Road, London SW7 2AZ, United Kingdom.