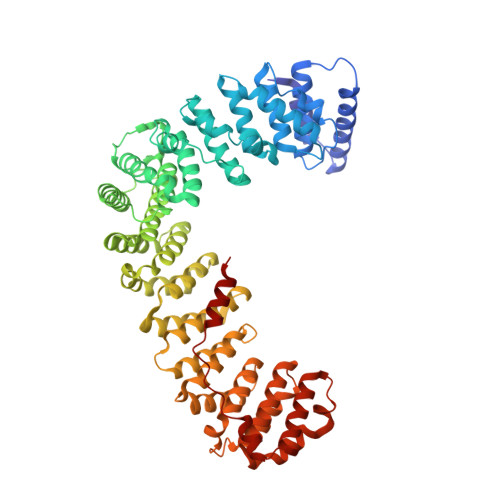
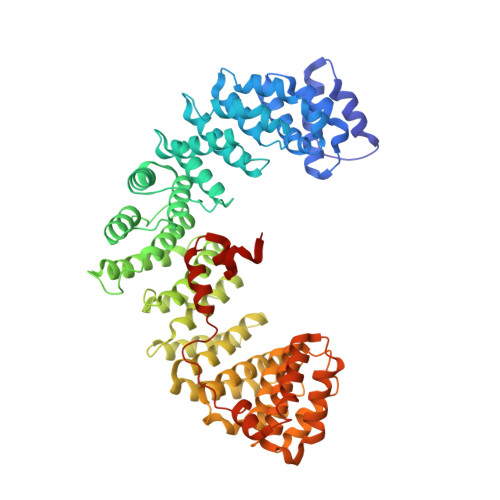
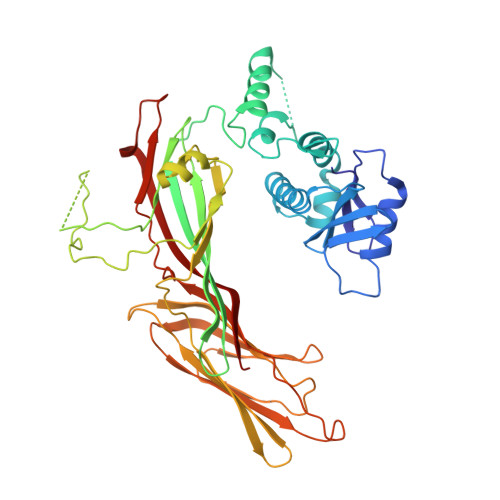
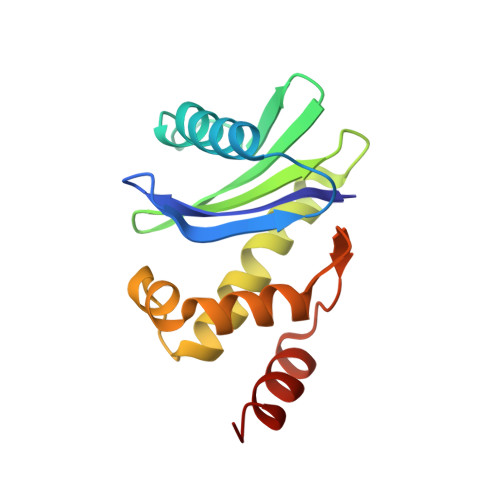
A Large Scale Conformational Change Couples Membrane Recruitment to Cargo Binding in the Ap2 Clathrin Adaptor Complex
Jackson, L.P., Kelly, B.T., Mccoy, A.J., Gaffry, T., James, L.C., Collins, B.M., Honing, S., Evans, P.R., Owen, D.J.(2010) Cell 141: 1241
- PubMed: 20603002
- DOI: https://doi.org/10.1016/j.cell.2010.05.006
- Primary Citation of Related Structures:
2XA7 - PubMed Abstract:
The AP2 adaptor complex (alpha, beta2, sigma2, and mu2 subunits) crosslinks the endocytic clathrin scaffold to PtdIns4,5P(2)-containing membranes and transmembrane protein cargo. In the "locked" cytosolic form, AP2's binding sites for the two endocytic motifs, YxxPhi on the C-terminal domain of mu2 (C-mu2) and [ED]xxxL[LI] on sigma2, are blocked by parts of beta2. Using protein crystallography, we show that AP2 undergoes a large conformational change in which C-mu2 relocates to an orthogonal face of the complex, simultaneously unblocking both cargo-binding sites; the previously unstructured mu2 linker becomes helical and binds back onto the complex. This structural rearrangement results in AP2's four PtdIns4,5P(2)- and two endocytic motif-binding sites becoming coplanar, facilitating their simultaneous interaction with PtdIns4,5P(2)/cargo-containing membranes. Using a range of biophysical techniques, we show that the endocytic cargo binding of AP2 is driven by its interaction with PtdIns4,5P(2)-containing membranes.
Organizational Affiliation:
Cambridge Institute for Medical Research, Department of Clinical Biochemistry, University of Cambridge, Hills Road, Cambridge CB2 0XY, UK.