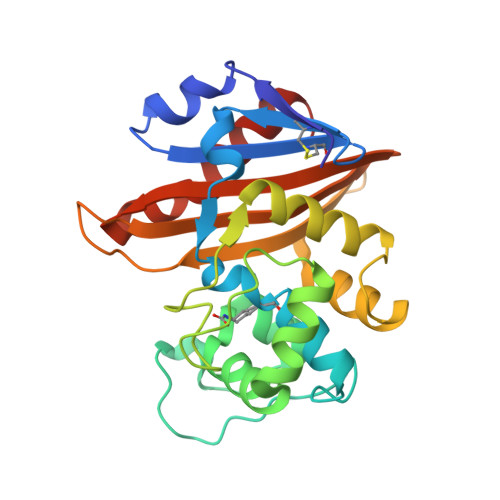
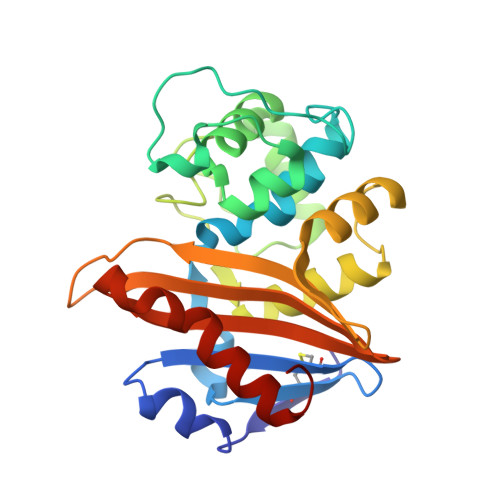
Three Factors that Modulate the Activity of Class D Beta-Lactamases and Interfere with the Post-Translational Carboxylation of Lys70.
Vercheval, L., Bauvois, C., Di Paolo, A., Borel, F., Ferrer, J.L., Sauvage, E., Matagne, A., Frere, J.M., Charlier, P., Galleni, M., Kerff, F.(2010) Biochem J 432: 495
- PubMed: 21108605
- DOI: https://doi.org/10.1042/BJ20101122
- Primary Citation of Related Structures:
2WGV, 2WGW, 2WKH, 2WKI - PubMed Abstract:
The activity of class D β-lactamases is dependent on Lys70 carboxylation in the active site. Structural, kinetic and affinity studies show that this post-translational modification can be affected by the presence of a poor substrate such as moxalactam but also by the V117T substitution. Val117 is a strictly conserved hydrophobic residue located in the active site. In addition, inhibition of class D β-lactamases by chloride ions is due to a competition between the side chain carboxylate of the modified Lys70 and chloride ions. Determination of the individual kinetic constants shows that the deacylation of the acyl-enzyme is the rate-limiting step for the wild-type OXA-10 β-lactamase.
Organizational Affiliation:
Laboratory of Biological Macromolecules, Centre for Protein Engineering, Institut de Chimie B6a, University of Liège, Liège 4000, Belgium.