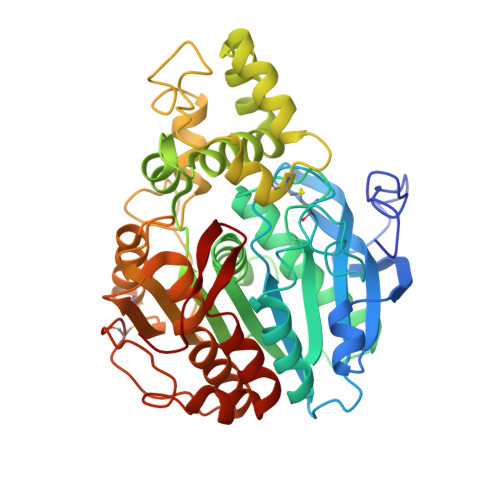
X-Ray Structure of Candida Antarctica Lipase a Shows a Novel Lid Structure and a Likely Mode of Interfacial Activation.
Ericsson, D.J., Kasrayan, A., Johansson, P., Bergfors, T., Sandstrom, A.G., Backvall, J.E., Mowbray, S.L.(2008) J Mol Biol 376: 109
- PubMed: 18155238
- DOI: https://doi.org/10.1016/j.jmb.2007.10.079
- Primary Citation of Related Structures:
2VEO - PubMed Abstract:
In nature, lipases (EC 3.1.1.3) catalyze the hydrolysis of triglycerides to form glycerol and fatty acids. Under the appropriate conditions, the reaction is reversible, and so biotechnological applications commonly make use of their capacity for esterification as well as for hydrolysis of a wide variety of compounds. In the present paper, we report the X-ray structure of lipase A from Candida antarctica, solved by single isomorphous replacement with anomalous scattering, and refined to 2.2-A resolution. The structure is the first from a novel family of lipases. Contrary to previous predictions, the fold includes a well-defined lid as well as a classic alpha/beta hydrolase domain. The catalytic triad is identified as Ser184, Asp334 and His366, which follow the sequential order considered to be characteristic of lipases; the serine lies within a typical nucleophilic elbow. Computer docking studies, as well as comparisons to related structures, place the carboxylate group of a fatty acid product near the serine nucleophile, with the long lipid tail closely following the path through the lid that is marked by a fortuitously bound molecule of polyethylene glycol. For an ester substrate to bind in an equivalent fashion, loop movements near Phe431 will be required, suggesting the primary focus of the conformational changes required for interfacial activation. Such movements will provide virtually unlimited access to solvent for the alcohol moiety of an ester substrate. The structure thus provides a basis for understanding the enzyme's preference for acyl moieties with long, straight tails, and for its highly promiscuous acceptance of widely different alcohol and amine moieties. An unconventional oxyanion hole is observed in the present structure, although the situation may change during interfacial activation.
Organizational Affiliation:
Department of Cell and Molecular Biology, Uppsala University, Biomedical Center, Box 596, SE-751 24 Uppsala, Sweden.