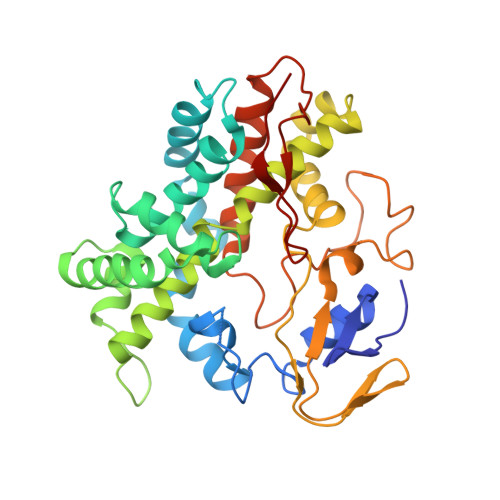
Crystal Structure and Properties of CYP231A2 from the Thermoacidophilic Archaeon Picrophilus torridus.
Ho, W.W., Li, H., Nishida, C.R., Ortiz de Montellano, P.R., Poulos, T.L.(2008) Biochemistry 47: 2071-2079
- PubMed: 18197710
- DOI: https://doi.org/10.1021/bi702240k
- Primary Citation of Related Structures:
2RFB, 2RFC - PubMed Abstract:
The crystal structure of a cytochrome P450 from the thermoacidophile Picrophilus torridus, CYP231A2 (PTO1399), has been solved. This structure reveals a wide open substrate access channel. To better understand ligand-induced structural transitions in CYP231A2, protein-ligand interactions were investigated using 4-phenylimidazole. Comparison of the ligand-free and -bound CYP231A2 structures shows conformational changes where the F and G helices swing as a single rigid body about a pivot point at the N-terminal end of the F helix, allowing the F helix region to dip toward the heme, resulting in closer contacts with the ligand. Thermal melting data illustrate that the melting temperature for CYP231A2 increases nearly 10 degrees C upon ligand binding, thus illustrating that the closed conformation is substantially more stable. Furthermore, spectroscopic data indicate that the active site is stable at pH 4.5, although, unusually, the thiolate ligand to the iron can be reversibly protonated. CYP231A2 does not exhibit structural features normally associated with thermophilic proteins such as an increase in salt bridge networks or extensive aromatic clustering. The increase in thermal stability instead is best correlated with the smaller size and shorter loops in CYP231A2 compared to other P450s.
Organizational Affiliation:
Departments of Molecular Biology and Biochemistry, Chemistry, and Pharmaceutical Sciences, University of California, Irvine, California 92697-3900, USA.