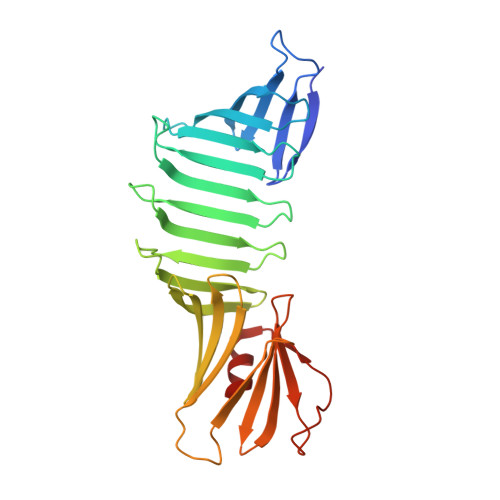
Aromatic cross-strand ladders control the structure and stability of beta-rich peptide self-assembly mimics
Biancalana, M., Makabe, K., Koide, A., Koide, S.(2008) J Mol Biol 383: 205-213
- PubMed: 18762191
- DOI: https://doi.org/10.1016/j.jmb.2008.08.031
- Primary Citation of Related Structures:
2OY7, 2OY8, 2OYB - PubMed Abstract:
Though beta-rich self-assemblies comprise a major structural class of polypeptides, a detailed understanding of the determinants of their structure and stability is lacking. In particular, the roles of repetitive stretches of side chains running the long axis of these beta-sheets, termed "cross-strand ladders," remain poorly characterized due to the inherently insoluble and heterogeneous nature of self-assemblies. To overcome these experimental challenges, we have established a complementary experimental system termed "peptide self-assembly mimics" (PSAMs). The PSAMs capture a defined number of self-assembly-like peptide repeats within a soluble beta-rich protein, making structural and energetic studies possible. In this work, we investigated the role of cross-strand ladders containing aromatic residues, which are prominent in self-assembling peptides. A combination of solution data and high-resolution crystal structures revealed that a single cross-strand ladder consisting solely of Tyr significantly stabilized, rigidified, and flattened the PSAM beta-sheet. These characteristics would stabilize each beta-sheet layer of a self-assembly and direct sheet conformations compatible with lamination. Our results therefore provide a rationale for the abundance of aromatic amino acids in fibril-forming peptides and establish important roles of cross-strand Tyr ladders in the structure and stability of beta-rich peptide self-assemblies.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, IL 60637, USA.