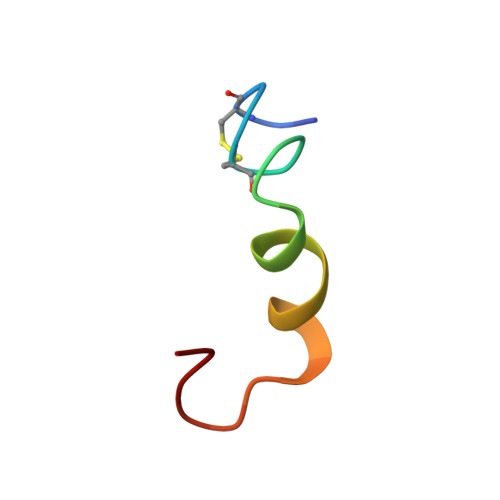
Segment TM7 from the cytoplasmic hemi-channel from V(O)-H(+)-V-ATPase includes a flexible region that has a potential role in proton translocation
Duarte, A.M., de Jong, E.R., Wechselberger, R., van Mierlo, C.P., Hemminga, M.A.(2007) Biochim Biophys Acta 1768: 2263-2270
- PubMed: 17573038
- DOI: https://doi.org/10.1016/j.bbamem.2007.05.014
- Primary Citation of Related Structures:
2NVJ - PubMed Abstract:
A 900-MHz NMR study is reported of peptide sMTM7 that mimics the cytoplasmic proton hemi-channel domain of the seventh transmembrane segment (TM7) from subunit a of H(+)-V-ATPase from Saccharomyces cerevisiae. The peptide encompasses the amino acid residues known to actively participate in proton translocation. In addition, peptide sMTM7 contains the amino acid residues that upon mutation cause V-ATPase to become resistant against the inhibitor bafilomycin. 2D TOCSY and NOESY (1)H-(1)H NMR spectra are obtained of sMTM7 dissolved in d(6)-DMSO and are used to calculate the three-dimensional structure of the peptide. The NMR-based structures and corresponding dynamical features of peptide sMTM7 show that sMTM7 is composed of two alpha-helical regions. These regions are separated by a flexible hinge of two residues. The hinge acts as a ball-and-joint socket and both helical segments move independently with respect to one another. This movement in TM7 is suggested to cause the opening and closing of the cytoplasmic proton hemi-channel and enables proton translocation.
Organizational Affiliation:
Laboratory of Biophysics, Wageningen University, Dreijenlaan 3, 6703 HA Wageningen, The Netherlands.