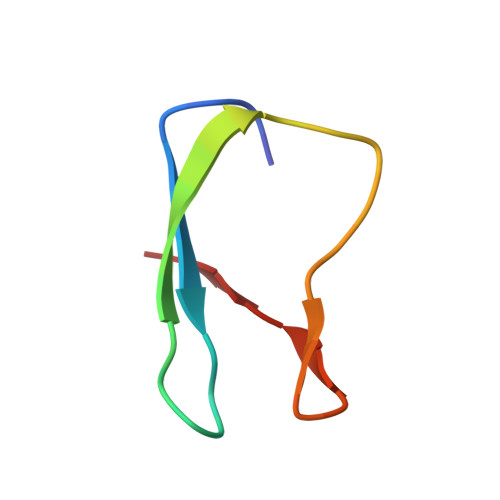
NMR determination of the global structure of the 113Cd derivative of desulforedoxin: investigation of the hydrogen bonding pattern at the metal center.
Goodfellow, B.J., Rusnak, F., Moura, I., Domke, T., Moura, J.J.(1998) Protein Sci 7: 928-937
- PubMed: 9568899
- DOI: https://doi.org/10.1002/pro.5560070410
- Primary Citation of Related Structures:
2LK6 - PubMed Abstract:
Desulforedoxin (Dx) is a simple homodimeric protein isolated from Desulfovibrio gigas (Dg) containing a distorted rubredoxin-like center with one iron coordinated by four cysteinyl residues (7.9 kDa with 36 amino acids per monomer). In order to probe the geometry and the H-bonding at the active site of Dx, the protein was reconstituted with 113Cd and the solution structure determined using 2D NMR methods. The structure of this derivative was initially compared with the NMR solution structure of the Zn form (Goodfellow BJ et al., 1996, J Biol Inorg Chem 1:341-353). Backbone amide protons for G4, D5, G13, L11 NH, and the Q14 NH side-chain protons, H-bonded in the X-ray structure, were readily exchanged with solvent. Chemical shift differences observed for amide protons near the metal center confirm the H-bonding pattern seen in the X-ray model (Archer M et al., 1995, J Mol Biol 251:690-702) and also suggest that H-bond lengths may vary between the Fe, Zn, and 113Cd forms. The H-bonding pattern was further probed using a heteronuclear spin echo difference (HSED) experiment; the results confirm the presence of NH-S H-bonds inferred from D2O exchange data and observed in the NMR family of structures. The presence of "H-bond mediated" coupling in Dx indicates that the NH-S H-bonds at the metal center have significant covalent character. The HSED experiment also identified an intermonomer "through space" coupling for one of the L26 methyl groups, indicating its proximity to the 113Cd center in the opposing monomer. This is the first example of an intermonomer "through space" coupling. Initial structure calculations produced subsets of NMR families with the S of C28 pointing away from or toward the L26 methyl: only the subset with the C28 sulfur pointing toward the L26 methyl could result in a "through space" coupling. The HSED result was therefore included in the structure calculations. Comparison of the Fe, Zn, and 113Cd forms of Dx suggests that the geometry of the metal center and the global fold of the protein does not vary to any great extent, although the H-bond network varies slightly when Cd is introduced. The similarity between the H-bonding pattern seen at the metal center in Dx, Rd (including H-bonded and through space-mediated coupling), and many zinc-finger proteins suggests that these H-bonds are structurally vital for stabilization of the metal centers in these proteins.
Organizational Affiliation:
Departamento de Química (and Centro de Química Fina e Biotecnologia), Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, Monte de Caparica, Portugal.