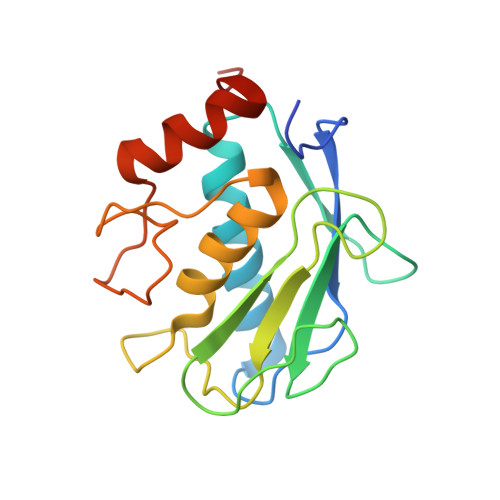
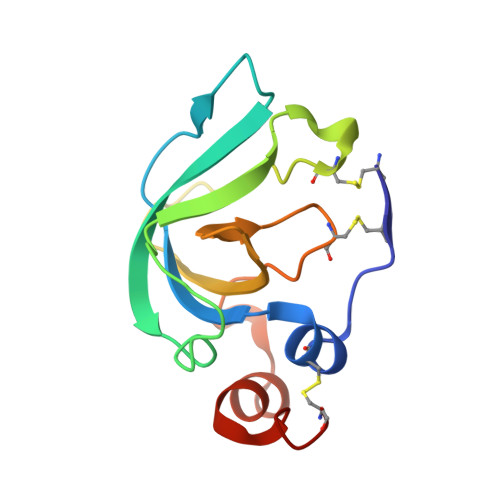
Crystal Structure of the Catalytic Domain of Matrix Metalloproteinase-1 in Complex with the Inhibitory Domain of Tissue Inhibitor of Metalloproteinase-1.
Iyer, S., Wei, S., Brew, K., Acharya, K.R.(2007) J Biol Chem 282: 364
- PubMed: 17050530
- DOI: https://doi.org/10.1074/jbc.M607625200
- Primary Citation of Related Structures:
2J0T - PubMed Abstract:
The mammalian collagenases are a subgroup of the matrix metalloproteinases (MMPs) that are uniquely able to cleave triple helical fibrillar collagens. Collagen breakdown is an essential part of extracellular matrix turnover in key physiological processes including morphogenesis and wound healing; however, unregulated collagenolysis is linked to important diseases such as arthritis and cancer. The tissue inhibitors of metalloproteinases (TIMPs) function in controlling the activity of MMPs, including collagenases. We report here the structure of a complex of the catalytic domain of fibroblast collagenase (MMP-1) with the N-terminal inhibitory domain of human TIMP-1 (N-TIMP-1) at 2.54 A resolution. Comparison with the previously reported structure of the TIMP-1/stromelysin-1 (MMP-3) complex shows that the mechanisms of inhibition of both MMPs are generally similar, yet there are significant differences in the protein-protein interfaces in the two complexes. Specifically, the loop between beta-strands A and B of TIMP-1 makes contact with MMP-3 but not with MMP-1, and there are marked differences in the roles of individual residues in the C-D connector of TIMP-1 in binding to the two MMPs. Structural rearrangements in the bound MMPs are also strikingly different. This is the first crystallographic structure that contains the truncated N-terminal domain of a TIMP, which shows only minor differences from the corresponding region of the full-length protein. Differences in the interactions in the two TIMP-1 complexes provide a structural explanation for the results of previous mutational studies and a basis for designing new N-TIMP-1 variants with restricted specificity.
Organizational Affiliation:
Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, UK.