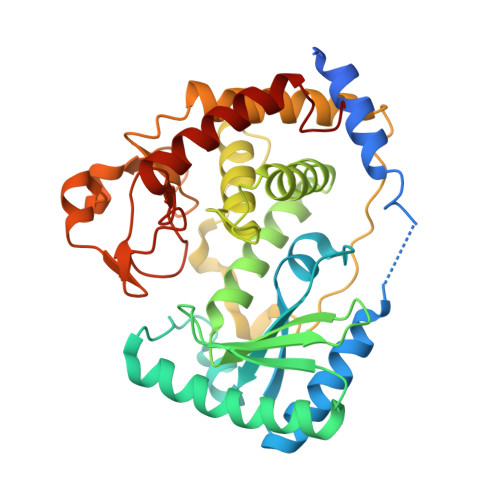
UTP-bound and Apo Structures of a Minimal RNA Uridylyltransferase.
Stagno, J., Aphasizheva, I., Rosengarth, A., Luecke, H., Aphasizhev, R.(2007) J Mol Biol 366: 882-899
- PubMed: 17189640
- DOI: https://doi.org/10.1016/j.jmb.2006.11.065
- Primary Citation of Related Structures:
2IKF, 2NOM - PubMed Abstract:
3'-Uridylylation of RNA is emerging as a phylogenetically widespread phenomenon involved in processing events as diverse as uridine insertion/deletion RNA editing in mitochondria of trypanosomes and small nuclear RNA (snRNA) maturation in humans. This reaction is catalyzed by terminal uridylyltransferases (TUTases), which are template-independent RNA nucleotidyltransferases that specifically recognize UTP and belong to a large enzyme superfamily typified by DNA polymerase beta. Multiple TUTases, recently identified in trypanosomes, as well as a U6 snRNA-specific TUTase enzyme in humans, are highly divergent at the protein sequence level. However, they all possess conserved catalytic and UTP recognition domains, often accompanied by various auxiliary modules present at the termini or between conserved domains. Here we report identification, structural and biochemical analyses of a novel trypanosomal TUTase, TbTUT4, which represents a minimal catalytically active RNA uridylyltransferase. The TbTUT4 consists of only two domains that define the catalytic center at the bottom of the nucleoside triphosphate and RNA substrate binding cleft. The 2.0 Angstroms crystal structure reveals two significantly different conformations of this TUTase: one molecule is in a relatively open apo conformation, whereas the other displays a more compact TUTase-UTP complex. A single nucleoside triphosphate is bound in the active site by a complex network of interactions between amino acid residues, a magnesium ion and highly ordered water molecules with the UTP's base, ribose and phosphate moieties. The structure-guided mutagenesis and cross-linking studies define the amino acids essential for catalysis, uracil base recognition, ribose binding and phosphate coordination by uridylyltransferases. In addition, the cluster of positively charged residues involved in RNA binding is identified. We also report a 2.4 Angstroms crystal structure of TbTUT4 with the bound 2' deoxyribonucleoside, which provides the structural basis of the enzyme's preference toward ribonucleotides.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697, USA.