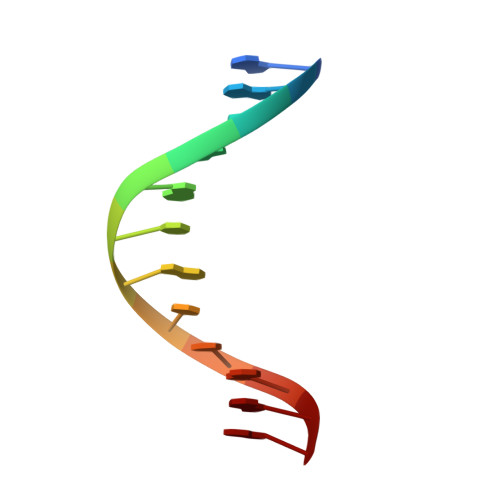
Base-displaced intercalated structure of the food mutagen 2-amino-3-methylimidazo[4,5-f]quinoline in the recognition sequence of the NarI restriction enzyme, a hotspot for -2 bp deletions.
Wang, F., DeMuro, N.E., Elmquist, C.E., Stover, J.S., Rizzo, C.J., Stone, M.P.(2006) J Am Chem Soc 128: 10085-10095
- PubMed: 16881637
- DOI: https://doi.org/10.1021/ja062004v
- Primary Citation of Related Structures:
2HKB, 2HKC - PubMed Abstract:
The solution structure of the oligodeoxynucleotide 5'-d(CTCGGCXCCATC)-3'.5'-d(GATGGCGCCGAG)-3' containing the heterocyclic amine 8-[(3-methyl-3H-imidazo[4,5-f]quinolin-2-yl)amino]-2'-deoxyguanosine adduct (IQ) at the third guanine in the NarI restriction sequence, a hot spot for -2 bp frameshifts, is reported. Molecular dynamics calculations restrained by distances derived from 24 (1)H NOEs between IQ and DNA, and torsion angles derived from (3)J couplings, yielded ensembles of structures in which the adducted guanine was displaced into the major groove with its glycosyl torsion angle in the syn conformation. One proton of its exocyclic amine was approximately 2.8 A from an oxygen of the 5' phosphodiester linkage, suggesting formation of a hydrogen bond. The carcinogen-guanine linkage was defined by torsion angles alpha' [N9-C8-N(IQ)-C2(IQ)] of 159 +/- 7 degrees and beta' [C8-N(IQ)-C2(IQ)-N3(IQ)] of -23 +/- 8 degrees . The complementary cytosine was also displaced into the major groove. This allowed IQ to intercalate between the flanking C.G base pairs. The disruption of Watson-Crick hydrogen bonding was corroborated by chemical-shift perturbations for base aromatic protons in the complementary strand opposite to the modified guanine. Chemical-shift perturbations were also observed for (31)P resonances corresponding to phosphodiester linkages flanking the adduct. The results confirmed that IQ adopted a base-displaced intercalated conformation in this sequence context but did not corroborate the formation of a hydrogen bond between the IQ quinoline nitrogen and the complementary dC [Elmquist, C. E.; Stover, J. S.; Wang, Z.; Rizzo, C. J. J. Am. Chem. Soc. 2004, 126, 11189-11201].
Organizational Affiliation:
Department of Chemistry, Center in Molecular Toxicology, Vanderbilt-Ingram Cancer Center, Vanderbilt University, Nashville, Tennessee 37235, USA.