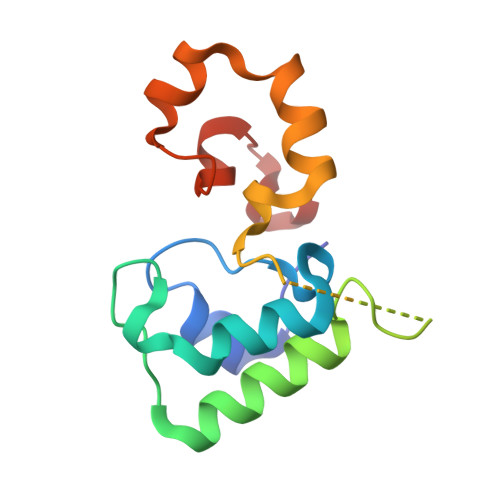
Structural and Functional Studies on DHC, the Diheme Cytochrome c from Rhodobacter sphaeroides, and Its Interaction with SHP, the sphaeroides Heme Protein
Gibson, H.R., Mowat, C.G., Miles, C.S., Li, B.R., Leys, D., Reid, G.A., Chapman, S.K.(2006) Biochemistry 45: 6363-6371
- PubMed: 16700547
- DOI: https://doi.org/10.1021/bi060288q
- Primary Citation of Related Structures:
2FW5, 2FWT - PubMed Abstract:
The diheme cytochrome c (DHC) from Rhodobacter sphaeroides is a soluble protein with a mass of 16 kDa that represents a new class of c-type cytochrome [Vandenberghe, I., et al. (1998) Biochemistry 37, 13075-13081]. The gene encoding DHC is associated with another encoding a cytochrome known as SHP (sphaeroides heme protein). It is believed that DHC is the electron donor for SHP, which is known to bind oxygen. To gain further insight into the properties and role of DHC, we have carried out structure-function studies on the protein and examined its interaction with SHP. The crystal structures of native and recombinant DHC have been determined to resolutions of 1.85 and 2.0 A, respectively. The structures show that DHC folds into two distinct domains each containing one heme. While the N-terminal domain is a class I cytochrome c, the C-terminal domain shows no similarity to any existing structures and thus constitutes a novel cytochrome c structural motif. The shortest, edge-to-edge, distance between the heme groups is 10.2 A, and this distance is bridged by Tyr31, thus ensuring fast internal electron transfer. DHC binds strongly to its proposed physiological partner, SHP (K(d) = 0.26 microM in 10 mM HEPES at pH 7.2 and 25 degrees C). However, at higher salt concentrations, the binding becomes much weaker, indicating the importance of electrostatic interactions. DHC is also very efficient in electron transfer to SHP with a second-order rate constant of 1.8 x 10(7) M(-)(1) s(-)(1) (at pH 7.2, 10 degrees C, and I = 500 mM). The reduction potentials of DHC and SHP are also suitably ordered for a favorable reaction with the hemes of DHC showing potentials of -310 and -240 mV, respectively, and that for SHP being -105 mV. These potentials are unaltered upon complex formation.
Organizational Affiliation:
School of Chemistry, University of Edinburgh, UK.