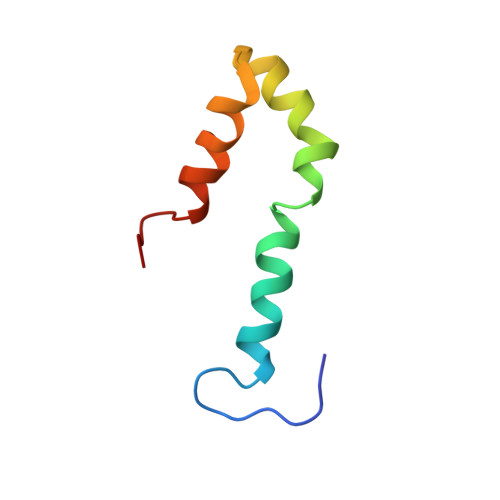
Solution structure of a protein denatured state and folding intermediate.
Religa, T.L., Markson, J.S., Mayor, U., Freund, S.M., Fersht, A.R.(2005) Nature 437: 1053-1056
- PubMed: 16222301
- DOI: https://doi.org/10.1038/nature04054
- Primary Citation of Related Structures:
1ZTR - PubMed Abstract:
The most controversial area in protein folding concerns its earliest stages. Questions such as whether there are genuine folding intermediates, and whether the events at the earliest stages are just rearrangements of the denatured state or progress from populated transition states, remain unresolved. The problem is that there is a lack of experimental high-resolution structural information about early folding intermediates and denatured states under conditions that favour folding because competent states spontaneously fold rapidly. Here we have solved directly the solution structure of a true denatured state by nuclear magnetic resonance under conditions that would normally favour folding, and directly studied its equilibrium and kinetic behaviour. We engineered a mutant of Drosophila melanogaster Engrailed homeodomain that folds and unfolds reversibly just by changing ionic strength. At high ionic strength, the mutant L16A is an ultra-fast folding native protein, just like the wild-type protein; however, at physiological ionic strength it is denatured. The denatured state is a well-ordered folding intermediate, poised to fold by docking helices and breaking some non-native interactions. It unfolds relatively progressively with increasingly denaturing conditions, and so superficially resembles a denatured state with properties that vary with conditions. Such ill-defined unfolding is a common feature of early folding intermediate states and accounts for why there are so many controversies about intermediates versus compact denatured states in protein folding.
Organizational Affiliation:
MRC Centre for Protein Engineering and Cambridge University Chemical Laboratories, MRC Centre, Hills Road, Cambridge CB2 2QH, UK.