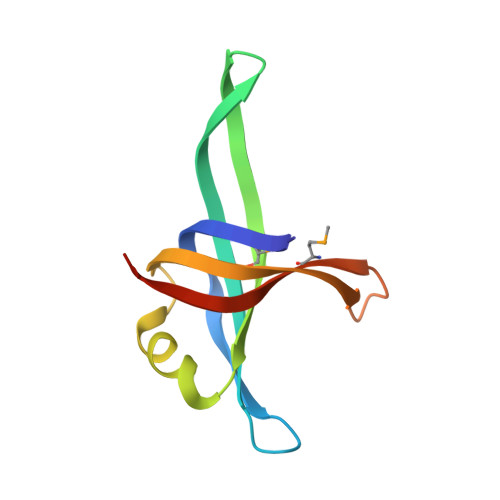
Crystal structure of a biologically functional form of PriB from Escherichia coli reveals a potential single-stranded DNA-binding site
Shioi, S., Ose, T., Maenaka, K., Shiroishi, M., Abe, Y., Kohda, D., Katayama, T., Ueda, T.(2005) Biochem Biophys Res Commun 326: 766-776
- PubMed: 15607735
- DOI: https://doi.org/10.1016/j.bbrc.2004.11.104
- Primary Citation of Related Structures:
1WOC - PubMed Abstract:
PriB is not only an essential protein necessary for the replication restart on the collapsed and disintegrated replication fork, but also an important protein for assembling of primosome onto PhiX174 genomic DNA during replication initiation. Here we report a 2.0-A-resolution X-ray structure of a biologically functional form of PriB from Escherichia coli. The crystal structure revealed that despite a low level of primary sequence identity, the PriB monomer, as well as the dimeric form, are structurally identical to the N-terminal DNA-binding domain of the single-stranded DNA-binding protein (SSB) from Escherichia coli, which possesses an oligonucleotides-binding-fold. The oligonucleotide-PriB complex model based on the oligonucleotides-SSB complex structure suggested that PriB had a DNA-binding pocket conserved in SSB from Escherichia coli and might bind to single-stranded DNA in the manner of SSB. Furthermore, surface plasmon resonance analysis and fluorescence measurements demonstrated that PriB binds single-stranded DNA with high affinity, by involving tryptophan residue. The significance of these results with respect to the functional role of PriB in the assembly of primosome is discussed.
Organizational Affiliation:
Department of Immunology, Graduate School of Pharmaceutical Sciences, Kyushu University, 3-1-1 Maidashi, Higashi-ku, Fukuoka 812-8582, Japan.