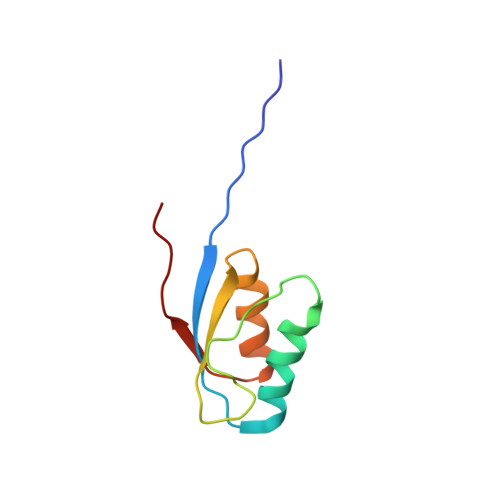
NMR Solution Structure of the Activation Domain of Human Procarboxypeptidase A2
Jimenez, M.A., Villegas, V., Santoro, J., Serrano, L., Vendrell, J., Aviles, F.X., Rico, M.(2003) Protein Sci 12: 296
- PubMed: 12538893
- DOI: https://doi.org/10.1110/ps.0227303
- Primary Citation of Related Structures:
1O6X - PubMed Abstract:
The activation domain of human procarboxypeptidase A2, ADA2h, is an 81-residue globular domain released during the proteolytic activation of the proenzyme. The role of this and other similar domains as assistants of the correct folding of the enzyme is not fully understood. The folding pathway of ADA2h was characterized previously, and it was also observed that under certain conditions it may convert into amyloid fibrils in vitro. To gain insight into these processes, a detailed description of its three-dimensional structure in aqueous solution is required so that eventual changes could be properly monitored. A complete assignment of the (1)H and (15)N resonances of ADA2h was performed, and the solution structure, as derived from a set of 1688 nonredundant constraints, is very well defined (pairwise backbone RMSD = 0.67 +/- 0.17 A for residues 10-80). The structure is composed of two antiparallel alpha-helices comprising residues 19-32 and 58-69 packed on the same side of a three-stranded beta-sheet spanning residues 10-15, 50-55, and 73-75. The global fold for the isolated human A2 activation domain is very similar to that of porcine carboxypeptidase B, as well as to the structure of the domain in the crystal of the intact human proenzyme. The observed structural differences relative to the intact human proenzyme are located at the interface between the activation domain and the enzyme and can be related with the activation mechanism. The backbone amide proton exchange behavior of ADA2h was also examined. The global free energy of unfolding obtained from exchange data of the most protected amide protons at pH 7.0 and 298K is 4.9 +/- 0.3 kcal.mole(-1), in good agreement with the values determined by thermal or denaturant unfolding studies.
Organizational Affiliation:
Instituto de Química-Física Rocasolano. C.S.I.C., Serrano, 119, 28006 Madrid, Spain.