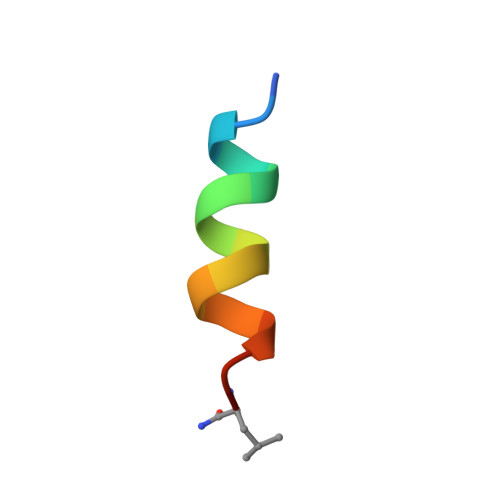
Interaction of mastoparan with membranes studied by 1H-NMR spectroscopy in detergent micelles and by solid-state 2H-NMR and 15N-NMR spectroscopy in oriented lipid bilayers.
Hori, Y., Demura, M., Iwadate, M., Ulrich, A.S., Niidome, T., Aoyagi, H., Asakura, T.(2001) Eur J Biochem 268: 302-309
- PubMed: 11168364
- DOI: https://doi.org/10.1046/j.1432-1033.2001.01880.x
- Primary Citation of Related Structures:
1D7N - PubMed Abstract:
Several complementary NMR approaches were used to study the interaction of mastoparan, a 14-residue peptide toxin from wasp venom, with lipid membranes. First, the 3D structure of mastoparan was determined using 1H-NMR spectroscopy in perdeuterated (SDS-d25) micelles. NOESY experiments and distance geometry calculations yielded a straight amphiphilic alpha-helix with high-order parameters, and the chemical shifts of the amide protons showed a characteristic periodicity of 3-4 residues. Secondly, solid-state 2H-NMR spectoscopy was used to describe the binding of mastoparan to lipid bilayers, composed of headgroup-deuterated dimyristoylglycerophosphocholine (DMPC-d4) and dimyristoylphosphatidylglycerol (DMPG). By correlating the deuterium quadrupole splittings of the alpha-segments and beta-segments, it was possible to differentiate the electrostatically induced structural response of the choline headgroup from dynamic effects induced by the peptide. A partial phase separation was observed, leading to a DMPG-rich phase and a DMPG-depleted phase, each containing some mastoparan. Finally, the insertion and orientation of a specifically 15N-labeled mastoparan (at position Ala10) in the bilayer environment was investigated by solid-state 15N-NMR spectroscopy, using macroscopically oriented samples. Two distinct orientational states were observed for the mastoparan helix, namely an in-plane and a trans-membrane alignment. The two populations of 90% in-plane and 10% trans-membrane helices are characterized by a mosaic spread of +/- 30 degrees and +/- 10 degrees, respectively. The biological activity of mastoparan is discussed in terms of a pore-forming model, as the peptide is known to be able to induce nonlamellar phases and facilitate a flip-flop between the monolayers.
Organizational Affiliation:
Department of Biotechnology, Tokyo University of Agriculture and Technology, Japan.