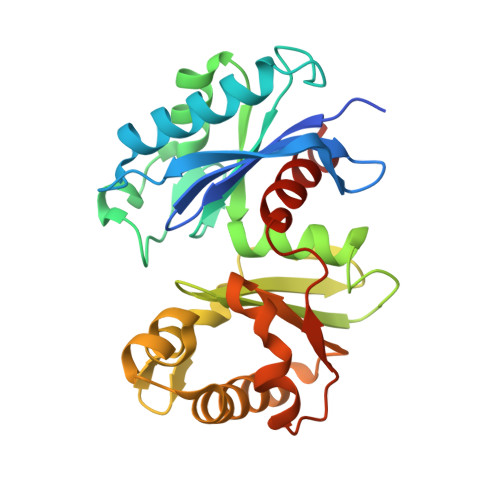
Crystal Structure of Bacterial Inorganic Polyphosphate/ATP-glucomannokinase: INSIGHTS INTO KINASE EVOLUTION
Mukai, T., Kawai, S., Mori, S., Mikami, B., Murata, K.(2004) J Biol Chem 279: 50591-50600
- PubMed: 15377666
- DOI: https://doi.org/10.1074/jbc.M408126200
- Primary Citation of Related Structures:
1WOQ - PubMed Abstract:
Inorganic polyphosphate (poly(P)) is a biological high energy compound presumed to be an ancient energy carrier preceding ATP. Several poly(P)-dependent kinases that use poly(P) as a phosphoryl donor are known to function in bacteria, but crystal structures of these kinases have not been solved. Here we present the crystal structure of bacterial poly(P)/ATP-glucomannokinase, belonging to Gram-positive bacterial glucokinase, complexed with 1 glucose molecule and 2 phosphate molecules at 1.8 A resolution, being the first among poly(P)-dependent kinases and bacterial glucokinases. The poly(P)/ATP-glucomannokinase structure enabled us to understand the structural relationship of bacterial glucokinase to eucaryotic hexokinase and ADP-glucokinase, which has remained a matter of debate. These comparisons also enabled us to propose putative binding sites for phosphoryl groups for ATP and especially for poly(P) and to obtain insights into the evolution of kinase, particularly from primordial poly(P)-specific to ubiquitous ATP-specific proteins.
Organizational Affiliation:
Department of Basic and Applied Molecular Biotechnology, Division of Food and Biological Science, Graduate School of Agriculture, Kyoto University, Uji, Kyoto 611-0011, Japan.