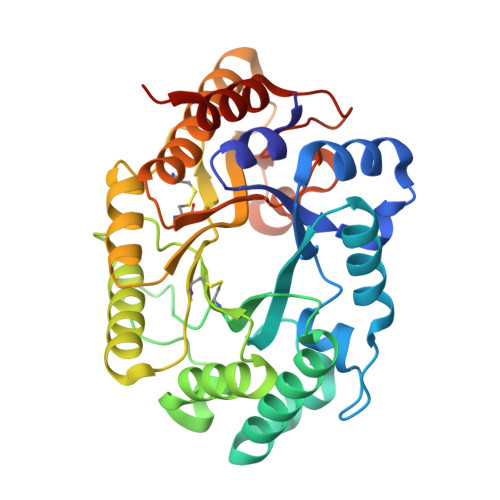
Structure and function of a family 10 beta-xylanase chimera of Streptomyces olivaceoviridis E-86 FXYN and Cellulomonas fimi Cex
Kaneko, S., Ichinose, H., Fujimoto, Z., Kuno, A., Yura, K., Go, M., Mizuno, H., Kusakabe, I., Kobayashi, H.(2004) J Biol Chem 279: 26619-26626
- PubMed: 15078885
- DOI: https://doi.org/10.1074/jbc.M308899200
- Primary Citation of Related Structures:
1V6Y - PubMed Abstract:
The catalytic domain of xylanases belonging to glycoside hydrolase family 10 (GH10) can be divided into 22 modules (M1 to M22; Sato, Y., Niimura, Y., Yura, K., and Go, M. (1999) Gene (Amst.) 238, 93-101). Inspection of the crystal structure of a GH10 xylanase from Streptomyces olivaceoviridis E-86 (SoXyn10A) revealed that the catalytic domain of GH10 xylanases can be dissected into two parts, an N-terminal larger region and C-terminal smaller region, by the substrate binding cleft, corresponding to the module border between M14 and M15. It has been suggested that the topology of the substrate binding clefts of GH10 xylanases are not conserved (Charnock, S. J., Spurway, T. D., Xie, H., Beylot, M. H., Virden, R., Warren, R. A. J., Hazlewood, G. P., and Gilbert, H. J. (1998) J. Biol. Chem. 273, 32187-32199). To facilitate a greater understanding of the structure-function relationship of the substrate binding cleft of GH10 xylanases, a chimeric xylanase between SoXyn10A and Xyn10A from Cellulomonas fimi (CfXyn10A) was constructed, and the topology of the hybrid substrate binding cleft established. At the three-dimensional level, SoXyn10A and CfXyn10A appear to possess 5 subsites, with the amino acid residues comprising subsites -3 to +1 being well conserved, although the +2 subsites are quite different. Biochemical analyses of the chimeric enzyme along with SoXyn10A and CfXyn10A indicated that differences in the structure of subsite +2 influence bond cleavage frequencies and the catalytic efficiency of xylooligosaccharide hydrolysis. The hybrid enzyme constructed in this study displays fascinating biochemistry, with an interesting combination of properties from the parent enzymes, resulting in a low production of xylose.
Organizational Affiliation:
National Food Research Institute, 2-1-12 Kannondai, Tsukuba, Ibaraki 305-8642, Japan. sakaneko@nfri.affrc.go.jp